Host: Noah Baker
Welcome back to the Nature Podcast. This week, we’ll be hearing about the placental microbiome…
Host: Nick Howe
And learning about advance in AI hardware. I’m Nick Howe.
Host: Noah Baker
And I’m Noah Baker.
[Jingle]
Interviewer: Nick Howe
You may think of the microbiome as the collection of microorganisms that live in our gut, and that would be right… to a point. But our gut isn’t the only place to host that party of microbes and for the record, the word ‘microbiome’ specifically refers to the genetic information of those microbes. But anyway, they don’t just live in our gut. They also live on our skin, our mouths, even in our eyes, but there is a part of the body where scientists can’t seem to agree on whether or not microbes dwell, and that is the placenta – the organ which grows during pregnancy to deal with the fetus’ vital functions. Here’s Gordon Smith, a researcher in maternal fetal medicine.
Interviewee: Gordon Smith
So, the sort of original view of the placenta has been that it was thought to be sterile and there was a paper a few years back in Science Translational Medicine which really sort of turned around the thinking on that, suggesting that there could be a placental microbiome – that is a population of bacteria that would normally be living in the placenta.
Interviewer: Nick Howe
If true, this could have important implications for the developing baby. Bacteria in the placenta could cause disease or be associated with complications in pregnancy. Alternatively, the placenta could be part of how the baby acquires its own microbiome. Since the Science Translational Medicine study, there have been various papers going back and forth questioning whether or not a placental microbiome exists. This week in Nature, Gordon throws his own hat into the ring with the biggest study so far, using samples from 537 British women. Gordon sequenced all of the material in the samples, discounted any human genetic material and then looked to see if any bacterial DNA was left. But it isn’t quite that simple.
Interviewee: Gordon Smith
One of the questions that we were aware of was how do you differentiate real signal that’s in the sample from contamination that might be caused by any number of different sources. So, the key thing that we did was to use two methods and also then to look at the level of agreement between the two methods.
Interviewer: Nick Howe
Gordon looked at fragmented DNA from his samples and also searched for specific bacterial genes. If he didn’t get the same result from both methods, he discounted the signals as contamination.
Interviewee: Gordon Smith
And we saw lots and lots of signals with each method but they didn’t agree, and what we’ve concluded is that the most likely reason they didn’t agree is that they were introduced through some form of contamination in the sort of laboratory analysis.
Interviewer: Nick Howe
Contamination was a real issue. For example, Gordon found cholera in his samples, but given that there hasn’t been a cholera outbreak in the UK for quite a while, it seemed unlikely it was in women’s placentas. And indeed, using the two-method approach, Gordon concluded that the cholera was more likely to have come from the sequencing facility itself. He also found evidence that some of the supposedly sterile reagents they were using were occasionally contaminated. After discounting suspected contamination, Gordon found that there was no bacteria present on the placenta in healthy pregnancies, but that’s not to say there wasn’t anything at all.
Interviewee: Gordon Smith
We found one real signal and that is a species of Streptococcus which is called group B strep.
Interviewer: Nick Howe
This bacterium was found in 5% of samples and Gordon and his team believe that they were due to infections during pregnancy rather than a microbiome. Altogether, they conclude there is no microbiome associated with the placenta. So that’s it, debate settled.
Interviewee: Kjersti Aagaard
Not at all.
Interviewer: Nick Howe
This is Kjersti Aagaard, a researcher in maternal fetal medicine who is the lead author on the Science Translational Medicine paper Gordon mentioned earlier.
Interviewee: Kjersti Aagaard
I actually think they are describing a placental microbiome.
Interviewer: Nick Howe
Kjersti believes that a lot of what Gordon is dismissing as contamination are actually examples of the placental microbiome. Gordon only considered a signal real if he detected the same exact species using both methods. Kjersti pointed out that it could be tricky to identify bacteria at a species level using these techniques. Also, some bacteria were dismissed as they were thought to have been acquired during vaginal birth, but Kjersti disagreed.
Interviewee: Kjersti Aagaard
We cannot shut down crucial lines of investigation that may make a real difference for women and their families, including their babies. I think there is a real signal here. I am thrilled and I applaud these investigators for the incredible work they’ve done, but I think we look at it through a different lens and we don’t necessarily need to disregard things as contaminate when we consider the biology and some fundamentally important technical considerations that led to some different conclusions.
Interviewer: Nick Howe
In fact, looking at Gordon’s data, Kjersti would conclude there is a placental microbiome. The difference, she says, is in the interpretation. So, the debate on whether or not the placental microbiome exists may not be settled, but studies like Gordon’s are still useful to understand how infections like the group B Streptococcus he found can occur during pregnancy. Here’s Gordon.
Interviewee: Gordon Smith
Infection of the baby with group B strep is the most common cause of death of the baby in the first weeks of life due to sepsis, and so that’s really our current area of study. What we’re now trying to do is to go through a large number of samples and see if the presence of this group B strep in the placenta carries any predictive association with complications for the baby.
Host: Nick Howe
That was Gordon Smith of Cambridge University here in the UK. You also heard from Kjersti Aagaard of the Baylor College of Medicine in the US. You can find Gordon’s paper over at nature.com, along with a News and Views article.
Host: Noah Baker
Later in the show, how HIV is becoming resistant to drugs – that’s in the News Chat. But now, it’s time for the Research Highlights, read this week by Shamini Bundell.
[Jingle]
Shamini Bundell
When it comes to the causes of allergies, there are a lot of possible culprits, from extremely clean houses to the overuse of antibiotics. Now, research has confirmed another factor – prescription antacids, taken to treat things like heartburn or stomach ulcers. A new analysis of over 8 million health records showed that people using prescription antacids were twice as likely to need anti-allergy medication in the following years. This may be because antacids prevent food being fully broken down in the stomach, meaning larger protein fragments can reach the intestines and potentially sensitise the immune system, leading to an allergic response. Digest more on this story at Nature Communications.
[Jingle]
Shamini Bundell
What’s the best way to find the source of a leak? In October 2017, labs across Europe noticed extremely high levels of a rare radioactive isotope in the atmosphere. The isotope – ruthenium-106 – wasn’t concentrated enough to be dangerous to people, but researchers were keen to figure out where it had come from. They traced the source to Russia, to approximately the area of a nuclear processing plant. At the time, Russian officials denied that the plant could be the source of the leak and instead blamed a disintegrating satellite. Now, a group of European researchers have examined exactly when the radiation was detected in different countries, and studied air movements at the time of detection. They narrowed down the source to a region in the Southern Urals where the nuclear plant is located. They concluded that the leak was likely caused by an unreported accident at the plant. You can trace that story back to its source in the Proceedings of the National Academy of Sciences.
[Jingle]
Host: Noah Baker
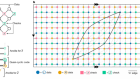
For many of us, artificial intelligence is already part of our lives. Neural networks – computer systems modelled on the human brain – have given rise to all sorts of applications, from face recognition to our interactions with personal assistants like Alexa or Google Home. And the rise of these technologies is showing no signs of slowing. But for this pace of change to continue, the hardware to run the neural networks may have to change. Much of what’s currently used wasn’t really designed for AI. Geoff Marsh caught up with Luke Fleet, one of Nature’s physics editors, to find out more and to discuss a new paper that’s tackling some of the problems facing the field.
Interviewee: Luke Fleet
As an editor, I get to see papers from lots of different fields, but I think AI hardware is my favourite topic and so I’m pretty excited about this new paper.
Interviewer: Geoff Marsh
Everyday there’s a sort of new AI technology that we can have on our phones – speech recognition, you know – that’s sort of software developments, isn’t it. You’re talking about hardware.
Interviewee: Luke Fleet
A lot of the AI computing that we do now is done on things like graphics cards and with CPUs which you would find on your laptop or on your PC, but they were developed for doing a different type of computation to what AI demands. It will probably be much more efficient and faster to do AI computing if we had hardware that was actually designed for AI computing. Often the AI computing is actually being done not on what we would call edge devices, so not on things like your mobile phone, they’re actually being done in data centres and these data centres often are not the most environmentally friendly or energy efficient places. So, there are quite scary stats around the amount of water that data centres would use, all the carbon emissions which potentially even rival something like the aviation industry or are projected to in the future if things don’t change.
Interviewer: Geoff Marsh
Wow, and when you say water, is that for sort of cooling?
Interviewee: Luke Fleet
Yeah, that’s one the problems, is that we have technology now that as you shrink down things like silicone transistors and put more and more on a chip, heating becomes a real, real problem. That’s actually one of the limiting factors behind how fast you can actually run CPUs – you can only run them at a certain speed because otherwise they’ll just melt.
Interviewer: Geoff Marsh
The reason I’ve dragged you down to the studio today then is to hear about a new development that’s come out of a Chinese research group, and they’ve created some new hardware I guess to solve some of these problems of inefficiency.
Interviewee: Luke Fleet
What they’ve tried to do is they’ve tried to build a hardware that allows people to actually run the types of neural networks that are currently operating in things like data centres, which are really software built neural networks, whilst at the same time allowing other types of networks, which we would normally see in specialised AI hardware chips. Some of the hardware that’s currently out there for AI computing, in what are called brain-inspired chips, often these are running a particular type of neural network, so these are called spiking neural networks, but interestingly these are quite different to the networks that we are currently interacting with and the way that you would programme these is also very, very different. What this team have managed to do is actually develop a chip that allows both of these different types to actually run on the same chip, and not just switching from one to the other – they actually kind of run them at the same time.
Interviewer: Geoff Marsh
And I’ve seen they’ve actually put out a video of the utility that they’ve put this new hybrid chip to and it’s an unmanned bicycle, and I have to say, it’s very eerie to watch. It’s essentially just a bicycle cycling itself and following a man and responding to his voice. Why have they chosen that as a demonstration and how impressive is it?
Interviewee: Luke Fleet
It’s a wonderful demonstration and it is quite impressive. I’m sure lots of people are aware of autonomous driving, though with autonomous bicycles, you have some different factors to think about. So, just getting the stability of a bike is not trivial, as I’m sure many toddlers will be able to tell you. And then there are many different inputs you can have with a bike. Obviously, there are obstacles on the road, but what they’ve tried to do is have voice commands as well and they’ve tried lots of different things to show that this bike can autonomously drive itself whilst constantly keeping its stability.
Interviewer: Geoff Marsh
And did it end up making it much more efficient? We were talking about the current AI tends to be really inefficient – has this kind of solved the problem?
Interviewee: Luke Fleet
So, that’s the really interesting and impressive thing about this paper, is that they’re demonstrating a chip that shows that you can use kind of some of the best bits from the neural network designs that are out there and potentially make AI computation a little bit more efficient. One of the problems is that if you then try to compare this type of chip’s performance to some of the other chips that are out there, it’s not necessarily clear that this chip can outperform the other chips. So, these more specialised chips that are running just one standard type of neural network, I’m sure that if you got one of these chips and tried to do similar experiments, you could probably do them. You might even be able to do them better than what this current chip can do.
Interviewer: Geoff Marsh
Why would you bother then trying to bring these two different sort of systems together in a hybrid like this?
Interviewee: Luke Fleet
The reason why the authors have tried to do this is that all of the AI we’ve talked about so far is really narrow AI – it’s trying to do a specific type of computation to optimise for a task. But what people really want to develop is what’s called AGI, so artificial general intelligence, so something that can go beyond these kind of specific optimisation tasks. And a good example for this is you can give a toddler a cartoon picture of a cat, and not even an actual picture but just a cartoon, and just from one image, it can go outside and recognise a cat, and the types of AI we have now are just so far away from that and I think everybody would agree that if you are to develop AGI then we are going to need hardware and new architectures. It’s not clear what these architectures are going to look like, but until people start to explore and start to innovate and start playing around with different designs, then it’s not clear actually how we’re going to progress with this. So, they’ve demonstrated in this paper that they’ve got this hybrid architecture that has lots of very clever tricks. It can be kind of reprogrammed, it’s multifunctional, it can run different types of networks, it can run different types of networks kind of alongside each other. It’s something that people weren’t necessarily thinking about doing. Part of the reason for that was that it’s not clear what advantage that gives you now, but as I say, the authors are trying to look beyond that, beyond narrow AI and say if we’re working towards artificial general intelligence, what hardware do we need to actually get that?
Host: Noah Baker
That was Luke Fleet, physics editor here at Nature. You can find out more about the chip that can cycle a bike in the paper – that’s at nature.com
Interviewer: Nick Howe
Finally on the show, it’s time for the News Chat and I’m joined in the studio by Flora Graham, editor of the Nature Briefing. Hello, Flora.
Interviewee: Flora Graham
Hello.
Interviewer: Nick Howe
Thanks for joining me. For our first story this week, the WHO has raised a red flag about resistance to HIV drugs. Flora, why has this alarm been raised?
Interviewee: Flora Graham
Well, the WHO has surveyed countries all around the world and they’ve found that 12 countries in Africa, Asia and the Americas are showing worrying levels of resistance to HIV drugs.
Interviewer: Nick Howe
So, what is a worrying level of resistance?
Interviewee: Flora Graham
Well, when more than 10% of adults with the virus have developed resistance, it’s not considered safe to give the same medication to others because it could actually increase resistance.
Interviewer: Nick Howe
Do we know why this resistance might be spreading?
Interviewee: Flora Graham
We’re not 100% sure why the resistance is spreading, but it could be because some people are interrupting their treatment, their treatment is starting and stopping, and there can be a lot of reasons for this. Pressure and stigma can all be factors in people’s lives that might cause them to stop treatment and then restart again.
Interviewer: Nick Howe
So, with this spread of resistance across these different countries, is there anything that is to be done?
Interviewee: Flora Graham
What the WHO recommends is to use a different HIV drug instead. It’s less susceptible to becoming drug-resistant and of course, being a completely new drug to the existing ones, it won’t be affected by this existing drug resistance.
Interviewer: Nick Howe
Well, we’ll have to see then if this new drug will be an effective treatment. For our second story, there’s news about human-animal hybrid experiments, so it appears that in Japan some new experiments have been approved. Flora, what exactly are the scientists trying to do here?
Interviewee: Flora Graham
So, these scientists, they want to create the very, very first step in what could ultimately become human organs inside animals that could be used for transplants in humans, but that’s very, very far down the road. We’re still talking about just the initial stages being approved.
Interviewer: Nick Howe
So, what are the ethical concerns with this work?
Interviewee: Flora Graham
Yes, this was banned in Japan until March. These kinds of human-animal hybrid embryos are controversial because we’re talking about human cells. We don’t know how they will affect the development of the animal. We don’t know ultimately what the endgame of this kind of research could possibly be. Some ethicists are concerned that we won’t be able to control how the human cells move in the animal body and they could even move beyond the targeted organ and we could end up with an animal’s brain that has human cells in it, and this raises kind of science-fiction level questions about human-animal hybrids and what they might ultimately become.
Interviewer: Nick Howe
So, are the researchers taking any steps to prevent this?
Interviewee: Flora Graham
Well, the researcher who is leading this project really emphasises that he’s starting in a very slow, targeted way and that actually, the animals will not be bought to term, so they’re talking about embryos here only at this point. The technique that they’re using is designed to make sure that any human cells are restricted to the organ that they’re trying to produce. So, for example, they actually create an animal embryo that actually lacks the gene necessary to develop a certain organ, like the pancreas in this case, and then they use human iPS cells, these are induced pluripotent stem cells that can in theory develop into any kind of cell. Then when the embryo develops, it uses those human iPS cells to develop that particular organ only.
Interviewer: Nick Howe
So, what are the next steps then for this research?
Interviewee: Flora Graham
Well, researchers are going to start right at the beginning growing human cells in mouse and rat embryos and then transplanting those embryos into surrogate animals. Eventually, way, way, way down the line, the goal is to produce animals with organs that are compatible with humans that can eventually be transplanted into people.
Interviewer: Nick Howe
Well, it sounds like there will be a few more hurdles to overcome before we get to organs through this technique. Thanks, Flora. Listeners, for more on those stories, head over to nature.com/news.
Host: Noah Baker
And that’s it for this week’s show. If you want to reach out to us about anything you’ve heard, or just want to say hi, then feel free to drop us a tweet (@NaturePodcast) or email us on podcast@nature.com. I’m Noah Baker.
Host: Nick Howe
And I’m Nick Howe. Thanks for listening.

 Read the paper: Human placenta has no microbiome but can contain potential pathogens
Read the paper: Human placenta has no microbiome but can contain potential pathogens
 Alarming surge in drug-resistant HIV uncovered
Alarming surge in drug-resistant HIV uncovered
 Japan approves first human-animal embryo experiments
Japan approves first human-animal embryo experiments