Abstract
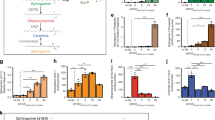
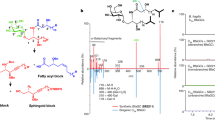
The essential nutrient choline is metabolized by gut bacteria to the disease-associated metabolite trimethylamine (TMA). However, most of the choline obtained via the diet and present in the human body is incorporated into larger metabolites, including the lipid phosphatidylcholine (PC). Here, we report that many choline-utilizing gut microorganisms can hydrolyse PC using a phospholipase D (PLD) enzyme and further convert the released choline to TMA. Genetic and in vitro characterization of the PLD from Escherichia coli MS 200-1 showed this enzyme is essential for bacterial hydrolysis of PC and prefers this substrate. PLDs are also found in gut bacterial isolates that are unable to convert choline to TMA, suggesting that additional members of the gut microbiota may influence access to this substrate. Unexpectedly, this PLD is only distantly related to characterized PLDs from pathogenic bacteria, suggesting a distinct evolutionary history. Together, these results reveal a previously underappreciated role for gut microorganisms in phospholipid metabolism and a potential target for inhibiting TMA production.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request.
References
Sender, R., Fuchs, S. & Milo, R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 14, e1002533-14 (2016).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Koppel, N. & Balskus, E. P. Exploring and understanding the biochemical diversity of the human microbiota. Cell Chem. Biol. 23, 18–30 (2016).
Arora, T. & Backhed, F. The gut microbiota and metabolic disease: current understanding and future perspectives. J. Intern. Med. 280, 339–349 (2016).
Kinross, J. M., Darzi, A. W. & Nicholson, J. K. Gut microbiome-host interactions in health and disease. Genome Med. 3, 14 (2011).
Zeisel, S. H. & da Costa, K.-A. Choline: an essential nutrient for public health. Nutr. Rev. 67, 615–623 (2009).
Zeisel, S. H. Choline: clinical nutrigenetic/nutrigenoic approaches for identification of functions and dietary requirements. World Rev. Nutr. Diet. 101, 73–83 (2010).
Hayward, H. R. & Stadtman, T. C. Anaerobic degradation of choline. I. Fermentation of choline by an anaerobic, cytochrome-producing bacterium, Vibrio cholinicus n. sp. J. Bacteriol. 78, 557–561 (1959).
Craciun, S. & Balskus, E. P. Microbial conversion of choline to trimethylamine requires a glycyl radical enzyme. Proc. Natl Acad. Sci. USA 109, 21307–21312 (2012).
Craciun, S., Marks, J. A. & Balskus, E. P. Characterization of choline trimethylamine-lyase expands the chemistry of glycyl radical enzymes. ACS Chem. Biol. 9, 1408–1413 (2014).
Krueger, S. K. & Williams, D. E. Mammalian flavin-containing monooxygenases: structure/function, genetic polymorphisms and role in drug metabolism. Pharmacol. Ther. 106, 357–387 (2005).
Mackay, R. J., McEntyre, C. J., Henderson, C., Lever, M. & George, P. M. Trimethylaminuria: causes and diagnosis of a socially distressing condition. Clin. Biochem. Rev. 32, 33–43 (2011).
Mitchell, S. C. & Smith, R. L. Trimethylaminuria: the fish malodor syndrome. Drug Metab. Dispos. 29, 517–521 (2001).
Sherriff, J. L., OSullivan, T. A., Properzi, C., Oddo, J. L. & Adams, L. A. Choline, its potential role in nonalcoholic fatty liver disease, and the case for human and bacterial genes. Adv. Nutr. 7, 5–13 (2016).
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63 (2012).
Tang, W. H. W. et al. Gut microbiota-dependent trimethylamine N-oxide (TMAO) pathway contributes to both development of renal insufficiency and mortality risk in chronic kidney disease. Circ. Res. 116, 448–455 (2015).
Miao, J. et al. Flavin-containing monooxygenase 3 as a potential player in diabetes-associated atherosclerosis. Nature 6, 6498 (2015).
Zeisel, S. H. & Warrier, M. Trimethylamine N-oxide, the microbiome, and heart and kidney disease. Annu. Rev. Nutr. 37, 157–181 (2017).
Corbin, K. D. & Zeisel, S. H. Choline metabolism provides novel insights into nonalcoholic fatty liver disease and its progression. Curr. Opin. Gastroen. 28, 159–165 (2012).
Jacobs, R. L., van der Veen, J. N. & Vance, D. E. Finding the balance: the role of S-adenosylmethionine and phosphatidylcholine metabolism in development of nonalcoholic fatty liver disease. Hepatology 58, 1207–1209 (2013).
Cohn, J. S., Kamili, A., Wat, E., Chung, R. W. S. & Tandy, S. Dietary phospholipids and intestinal cholesterol absorption. Nutrients 2, 116–127 (2010).
Zeisel, S. H., Mar, M.-H., Howe, J. C. & Holden, J. M. Concentrations of choline-containing compounds and betaine in common foods. J. Nutr. 133, 1302–1307 (2003).
Northfield, T. C. & Hofmann, A. F. Biliary lipid output during three meals and an overnight fast. I. Relationship to bile acid pool size and cholesterol saturation of bile in gallstone and control subjects. Gut 16, 1–11 (1975).
Braun, A. et al. Alterations of phospholipid concentration and species composition of the intestinal mucus barrier in ulcerative colitis: a clue to pathogenesis. Inflam. Bowel Dis. 15, 1705–1720 (2009).
Tang, W. H. W. et al. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N. Engl. J. Med. 368, 1575–1584 (2013).
Romano, K. A., Vivas, E. I., Amador-Noguez, D. & Rey, F. E. Intestinal microbiota composition modulates choline bioavailability from diet and accumulation of the proatherogenic metabolite trimethylamine-N-oxide. mBio 6, e02481-14 (2015).
Romano, K. A. et al. Metabolic, epigenetic, and transgenerational effects of gut bacterial choline consumption. Cell Host Microb. 22, 279–290 (2017).
Heller, M. Phospholipase D. Adv. Lipid Res. 16, 267–326 (1978).
Waite, M. The PLD superfamily: insights into catalysis. Biochim. Biophys. Acta 1439, 187–197 (1999).
Wang, X. Multiple forms of phospholipase D in plants: the gene family, catalytic and regulatory properties, and cellular functions. Prog. Lipid Res. 39, 109–149 (2000).
Frohman, M. A., Sung, T. C. & Morris, A. J. Mammalian phospholipase D structure and regulation. Biochim. Biophys. Acta 1439, 175–186 (1999).
Lery, L. M. S. et al. Comparative analysis of Klebsiella pneumoniae genomes identifies a phospholipase D family protein as a novel virulence factor. BMC Biol. 12, 41 (2014).
Uesugi, Y. & Hatanaka, T. Phospholipase D mechanism using Streptomyces PLD. BBA-Mol. Cell Biol. L. 1791, 962–969 (2009).
Barksdale, L., Linder, R., Sulea, I. T. & Pollice, M. Phospholipase D activity of Corynebacterium pseudotuberculosis (Corynebacterium ovis) and Corynebacterium ulcerans, a distinctive marker within the genus Corynebacterium. J. Clin. Microbiol. 13, 335–343 (1981).
Jacobs, A. C. et al. Inactivation of phospholipase D diminishes Acinetobacter baumannii pathogenesis. Infect. Immun. 78, 1952–1962 (2010).
Sung, T. C. et al. Mutagenesis of phospholipase D defines a superfamily including a trans-Golgi viral protein required for poxvirus pathogenicity. EMBO J. 16, 4519–4530 (1997).
Leiros, I., McSweeney, S. & Hough, E. The reaction mechanism of phospholipase D from Streptomyces sp. strain PMF. Snapshots along the reaction pathway reveal a pentacoordinate reaction intermediate and an unexpected final product. J. Mol. Biol. 339, 805–820 (2004).
Spencer, C. & Brown, H. A. Biochemical characterization of a Pseudomonas aeruginosa phospholipase D. Biochemistry 54, 1208–1218 (2015).
Liscovitch, M., Czarny, M., Fiucci, G. & Tang, X. Phospholipase D: molecular and cell biology of a novel gene family. Biochem. J. 345, 401–415 (2000).
Gottlin, E. B., Rudolph, A. E., Zhao, Y., Matthews, H. R. & Dixon, J. E. Catalytic mechanism of the phospholipase D superfamily proceeds via a covalent phosphohistidine intermediate. Proc. Natl Acad. Sci. USA 95, 9202–9207 (1998).
Yang, H. & Roberts, M. F. Cloning, overexpression, and characterization of a bacterial Ca2+-dependent phospholipase D. Protein Sci. 11, 2958–2968 (2009).
Bodea, S., Funk, M. A., Balskus, E. P. & Drennan, C. L. Molecular basis of C–N bond cleavage by the glycyl radical enzyme choline tTrimethylamine-lyase. Cell Chem. Biol. 23, 1206–1216 (2016).
Su, W. et al. 5-Fluoro-2-indolyl des-chlorohalopemide (FIPI), a phospholipase D pharmacological inhibitor that alters cell spreading and inhibits chemotaxis. Mol. Pharmacol. 75, 437–446 (2009).
Scott, S. A. et al. Discovery of desketoraloxifene analogues as inhibitors of mammalian, Pseudomonas aeruginosa, and NAPE phospholipase D enzymes. ACS Chem. Biol. 10, 421–432 (2015).
McNamara, P. J., Bradley, G. A. & Songer, J. G. Targeted mutagenesis of the phospholipase D gene results in decreased virulence of Corynebacterium pseudotuberculosis. Mol. Microbiol. 12, 921–930 (1994).
Hinnebusch, B. J. et al. Role of Yersinia murine toxin in survival of Yersinia pestis in the midgut of the flea vector. Science 296, 733–735 (2002).
Edwards, J. L. & Apicella, M. A. Neisseria gonorrhoeae PLD directly interacts with Akt kinase upon infection of primary, human, cervical epithelial cells. Cell Microbiol. 8, 1253–1271 (2006).
Dawson, R. M. Phosphorylcholine in rat tissues. Biochem. J. 60, 325–328 (1955).
Stremmel, W. Retarded release phosphatidylcholine benefits patients with chronic active ulcerative colitis. Gut 54, 966–971 (2005).
Karner, M. et al. First multicenter study of modified release phosphatidylcholine ‘LT-02’ in ulcerative colitis: a randomized, placebo-controlled trial in mesalazine-refractory courses. Am. J. Gastroenterol. 109, 1041–1051 (2014).
Driskell, L. O. et al. Directed mutagenesis of the Rickettsia prowazekii pldg gene encoding phospholipase D. Infect. Immun. 77, 3244–3248 (2009).
Zhou, M., Diwu, Z., Panchuk-Voloshina, N. & Haugland, R. P. A stable nonfluorescent derivative of resorufin for the fluorometric determination of trace hydrogen peroxide: applications in detecting the activity of phagocyte NADPH oxidase and other oxidases. Anal. Biochem. 253, 162–168 (1997).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Sievers, F. et al. Fast, scalable generation of high‐quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Reynolds, S. M., Kall, L., Riffle, M. E., Bilmes, J. A. & Noble, W. S. Transmembrane topology and signal peptide prediction using dynamic bayesian networks. PLoS Comput. Biol. 4, e10002123 (2008).
Tsirigos, K. D., Peters, C., Shu, N., Kall, L. & Elofsson, A. The TOPCONS web server for combined membrane protein topology and signal peptide prediction. Nucleic Acids Res. 43, W401–W407 (2015).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430, 2237–2243 (2018).
Jiang, F. et al. A Pseudomonas aeruginosa type VI secretion phospholipase D effector targets both prokaryotic and eukaryotic cells. Cell Host Microbe 15, 600–610 (2014).
Letunic, I. & Bork, P. Interactive tree of life (iTOL)v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Acknowledgements
The authors thank L. Rajakovich, B. Schneider, S. Irwin, N. Braffman and M. McCallum for helpful discussions and comments on the manuscript. The authors also thank K. Romano and F. Rey for helpful discussions, the provision of strains, and experimental design suggestions. The authors acknowledge financial support from the National Science Foundation (IDBR TYPE A DBI-1832846, EAGER MCB-1650086), the Camille Dreyfus Teacher-Scholar Award (TC-15-003), a Packard Fellowship for Science and Engineering (2013-39267), and a Blavatnik Biomedical Accelerator grant.
Author information
Authors and Affiliations
Contributions
C.L.C. and E.P.B. conceived the study. C.L.C and A.M.d.C. conceived and performed genetic manipulation experiments and analysed the data. C.L.C conceived and performed all biochemical, bioinformatic, and co-culturing experiments and analysed the data. C.L.C. and E.P.B. wrote the manuscript. C.L.C., A.M.d.C. and E.P.B. edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–6, Supplementary Figures 1–8.
Rights and permissions
About this article
Cite this article
Chittim, C.L., Martínez del Campo, A. & Balskus, E.P. Gut bacterial phospholipase Ds support disease-associated metabolism by generating choline. Nat Microbiol 4, 155–163 (2019). https://doi.org/10.1038/s41564-018-0294-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0294-4
This article is cited by
-
The Role of Gut Microbiota and Trimethylamine N-oxide in Cardiovascular Diseases
Journal of Cardiovascular Translational Research (2023)
-
Development of a thiostrepton-free system for stable production of PLD in Streptomyces lividans SBT5
Microbial Cell Factories (2022)
-
Calorie restriction improves metabolic state independently of gut microbiome composition: a randomized dietary intervention trial
Genome Medicine (2022)
-
Effect of microbiota metabolites on the progression of chronic hepatitis B virus infection
Hepatology International (2021)
-
Changes of intestinal microflora of breast cancer in premenopausal women
European Journal of Clinical Microbiology & Infectious Diseases (2021)