Abstract
Aneuploidy has been recognized as a hallmark of tumorigenesis for more than 100 years, but the connection between chromosomal errors and malignant growth has remained obscure. New evidence emerging from both basic and clinical research has illuminated a complicated relationship: despite its frequency in human tumours, aneuploidy is not a universal driver of cancer development and instead can exert substantial tumour-suppressive effects. The specific consequences of aneuploidy are highly context dependent and are influenced by a cell’s genetic and environmental milieu. In this Review, we discuss the diverse facets of cancer biology that are shaped by aneuploidy, including metastasis, drug resistance and immune recognition, and we highlight aneuploidy’s distinct roles as both a tumour promoter and an anticancer vulnerability.
Similar content being viewed by others
Introduction
The term ‘aneuploidy’ describes a state in which a cell’s karyotype is not a whole-number multiple of an organism’s normal haploid complement. In humans, aneuploidy is the most common cause of miscarriages and intellectual disabilities1,2. Aneuploidy is also a hallmark of malignant growth: approximately 90% of tumours display some degree of whole-chromosome aneuploidy3. Yet, recent evidence has demonstrated that aneuploidy can have both oncogenic and tumour-suppressive functions. By studying aneuploidy in genetically defined cell lines and mice, we have learned that the outcomes of aneuploidy can be shaped by tissue type, the identity of the affected chromosome and the manner in which an aneuploidy arises. Here, we highlight new research on the consequences of aneuploidy and its ability to influence multiple aspects of tumour development.
To clarify the terms used in this Review, we specifically use ‘aneuploidy’ to describe an imbalanced karyotype. If a cell gains an equal set of all chromosomes, then that ‘balanced’ state can be described as polyploidy, a condition that has been recently reviewed elsewhere4,5,6. On a fundamental level, ‘aneuploidy’ can also refer to a spectrum of related genetic alterations: a subchromosomal or segmental duplication that affects the copy number of a few genes still results in an aneuploidy-like genomic imbalance7,8. In this Review, we focus primarily on arm-length and whole-chromosome alterations that create imbalances in hundreds or thousands of genes at once, although we recognize that the consequences of segmental copy number alterations may closely resemble aneuploidy-like pathologies.
An overview of aneuploidy in cancer
In the late nineteenth century, the German scientist David von Hansemann described the appearance of abnormal mitotic figures in tissue sections from human tumours9. Subsequently, Theodor Boveri suggested that these chromosomal errors were actually the root cause of cancer, writing “a malignant tumour cell is … a cell with a specific abnormal chromosome constitution”10. A century of additional research into molecular oncology has complicated Boveri’s hypothesis. On a fundamental level, Boveri is incorrect: we now know that cancers arise due to mutations in oncogenes and tumour suppressors, and no karyotype has yet been discovered that is sufficient to transform a cell. At the same time, we have also learned that aneuploidy is exceptionally widespread. In some cancers, specific aneuploidies are observed more frequently than mutations in any known oncogene or tumour suppressor. For instance, approximately 60% of breast cancers harbour a trisomy of chromosome arm 1q11, making this alteration more common than mutations in PIK3CA (39% of breast tumours), TP53 (34%) or any other single gene10. Overall, about 97% of breast cancers exhibit one or more aneuploidies11.
Within individual cancer types, the frequency of aneuploidy can range from ~25% (thyroid carcinomas) to nearly 100% (glioblastomas)11. Some cancers display characteristic aneuploidy patterns: most glioblastomas gain an extra copy of chromosome 7 and lose a copy of chromosome 10, for instance. Other cancers display a very wide range of aneuploidies, which can include both clonal and subclonal alterations12. Indeed, aneuploidy is a major source of intratumour heterogeneity in many different cancers13.
Aneuploidy frequently arises early in tumorigenesis: single-chromosome gains and losses are often found in polyps, adenomas and other precancerous lesions14,15,16,17,18,19. However, sequencing studies have also revealed that certain chromosomal alterations, such as the gain of chromosome 7 in lung adenocarcinomas, tend to occur late in tumour development, potentially after cells have acquired prior mutations that allow them to tolerate the effects of aneuploidy20. Chromosome gains and losses typically result in a proportional change in the expression of genes on an affected chromosome, leading to an increase in gene expression from trisomic chromosomes and a decrease in expression from monosomies21,22,23,24,25,26,27,28. Finally, while the exact causes of aneuploidy remain obscure, some aneuploidies may arise from a phenomenon known as chromosomal instability (CIN), a state in which cells frequently mis-segregate whole chromosomes. It is important to note that CIN and aneuploidy are related but not identical: aneuploidy describes an imbalanced karyotype, while CIN describes a condition that accelerates the development of aneuploidy29.
Historically, aneuploidy has been resistant to close analysis. While single oncogenes and tumour suppressors have been studied for several decades using the standard tools of molecular genetics, manipulating whole chromosomes has been technically challenging. Thus, much of what we know comes from observational studies, in which particular aneuploidies have been observed to occur in certain diseases or in association with specific phenotypes. Yet, the aneuploidy that develops organically during tumorigenesis is caused by or jointly arises with other mutations, confounding our ability to understand cause and effect. To overcome these obstacles, researchers have developed a variety of approaches to model aneuploidy in cell lines and mice. Importantly, these approaches generate karyotypic diversity in otherwise-identical genetic backgrounds, thereby allowing researchers to isolate the effects of specific aneuploidies on tumorigenesis, absent any confounding genetic alterations.
In this Review, we focus on the relationship between aneuploidy and various malignant processes, including aneuploidy’s dual role as a cancer driver and a potential therapeutic vulnerability. Additionally, we point readers to several excellent reviews on related topics, including cellular stresses that are caused by aneuploidy30, the role of CIN in tumorigenesis31 and the interaction between tissue identity and aneuploidy13.
Modelling aneuploidy
Several methods have been used to engineer aneuploid mammalian cells (Fig. 1a,b; Table 1). In one approach, microcell-mediated chromosome transfer has been used to deliver specific donor chromosomes into recipient diploid cell lines32,33. Microcell-mediated chromosome transfer thereby creates defined human trisomies that can subsequently be compared with the parental cell line or with trisomies of other chromosomes. In another approach, diploid cells are transiently treated with small-molecule inhibitors of the spindle-assembly checkpoint kinase MPS1 (also known as TTK)34. Exposure to an MPS1 inhibitor generates a period of CIN, creating a pool of random aneuploidies that can be studied in bulk or isolated through single-cell cloning. Finally, Cre–lox-mediated recombination coupled with the expression of drug-resistance markers has been used to select for aneuploid cells in culture35.
a | Microcell-mediated chromosome transfer can be used to generate cell lines that have gained one or more chromosomes. b | Inhibitors of the spindle-assembly checkpoint (SAC) kinase MPS1 can be used to generate populations of aneuploid cells. These heterogeneous populations can be studied directly or single-cell cloning can be used to isolate clones with defined karyotypes. c–e | Chromosomal instability (CIN) mouse models have been generated by introducing mutations that eliminate kinetochore components, attenuate the SAC or stabilize microtubule–kinetochore attachments (part c), by introducing mutations that cause centrosome amplification (part d) and by introducing mutations in the cohesin complex that interfere with chromosome cohesion (part e). APC/C, anaphase-promoting complex/cyclosome.
To complement these cell line models, researchers have also sought to elucidate the consequences of aneuploidy in an in vivo setting. However, all autosomal trisomies and monosomies in the mouse are embryonically lethal36,37. To circumvent this limitation, aneuploid embryos can be generated by crossing mice that harbour naturally occurring Robertsonian translocations, and then aneuploid fibroblasts and various stem cell populations can be isolated, expanded in culture and analysed32,36,38,39. These cells have also been used to generate aneuploid–euploid chimaeras, either by mixing aneuploid and euploid embryonic stem cells40,41 or by reconstituting the blood supply of wild-type mice with aneuploid haematopoietic stem cells39,42. Finally, researchers have developed several mouse models of Down syndrome (caused by a trisomy of chromosome 21 in humans), which are described in more detail later43,44,45.
While constitutive aneuploidies are generally lethal, the effects of aneuploidy can also be studied by genetically engineering mice to induce somatic CIN. Approaches that have been described include attenuating the spindle-assembly checkpoint46, eliminating kinetochore components47,48, stabilizing microtubule–kinetochore attachments49,50, inducing centrosome amplification51,52 and disrupting chromosomal cohesion53,54 (Fig. 1c–e). Cells from these models harbour various degrees of mosaic aneuploidy in different tissues throughout their bodies and have been used to investigate the relationship between chromosome segregation errors and tumorigenesis. Several CIN models are summarized in Table 2, and their implications for cancer are discussed herein.
The role of aneuploidy in cancer
Many aneuploidies are capable of functioning as tumour suppressors
Despite the ubiquity of aneuploidy in cancer, engineered aneuploid cells typically exhibit decreased fitness and tumorigenicity27,32,38. Consistent with earlier results obtained in yeast55, human and mouse cells that carry a single extra chromosome exhibit a pronounced cell cycle delay, proteotoxic stress, metabolic alterations and genomic instability24,25,33,56,57,58. Aneuploid stem cells are outcompeted by euploid stem cells in mosaic embryos and when co-injected to reconstitute the mouse haematopoietic system39,40,41. Finally, aneuploid cells are resistant to transformation with oncogenes, senesce prematurely and grow poorly as tumour xenografts32,59. In total, these results suggest that aneuploidy is in fact capable of functioning as a tumour suppressor (Fig. 2).
a | Most aneuploidies display antitumorigenic properties that frequently lead to cell death or senescence. Rarely, aneuploid karyotypes arise that exhibit cancer-favouring properties, including immune evasion, drug resistance and oncogene overexpression. These aneuploidies can be selected over time, thereby increasing in abundance. b | The level of chromosomal instability (CIN) influences tumorigenic potential: very high rates of CIN lead to cell death, but lower levels of CIN can be tolerated and produce favourable karyotypes. However, the ‘ideal’ degree of CIN is influenced by tissue identity. c | CIN leads to heterogeneous aneuploidy, which can lead to either tumour-inhibitory effects or tumour progression. Senescent cells can contribute to both tumour inhibition and tumour progression through mechanisms that are not yet fully understood. d | Diagram depicting how Down syndrome simultaneously leads to increased haematological cancer risk, and decreased cancer risk in most solid tissues. SASP, senescence-associated secretory phenotype.
These observations have led to the suggestion that certain genetic alterations are required to ‘tolerate’ the deleterious effects of aneuploidy. In budding yeast, aneuploid cells spontaneously acquire mutations in the ubiquitin–proteasome pathway that enhance their proliferative capacity60. In mammalian cells, loss of the tumour suppressor p53 has been reported to facilitate the growth of aneuploid cells61,62,63,64. However, careful comparisons have highlighted that the loss of p53 enhances both euploid cell and aneuploid cell proliferation, and blocking p53 signalling is insufficient to fully restore aneuploid cell fitness32. Other proposed mechanisms for aneuploidy tolerance include loss of the proapoptotic protein BCL-9L and the stress kinase p38α (also known as MAPK14)65,66, although the relative paucity of mutations in the genes encoding these proteins in human tumours suggests they may not be common mechanisms for aneuploidy tolerance. In general, how cancers are able exhibit robust growth while harbouring highly aneuploid karyotypes remains incompletely understood.
Aneuploidy can promote growth under suboptimal conditions
While aneuploidy is deleterious for cell fitness in most cases, it has also been found to provide a selective advantage in certain challenging environments. For instance, while cancer cells harbouring single extra chromosomes grow poorly under standard culture conditions, the same trisomies were found to confer a proliferative advantage when those cells were cultured in hypoxic or serum-free conditions67. Similarly, two recent preprints demonstrate that the induction of CIN can allow cancer cells to adapt and survive in the presence of diverse anticancer agents68,69, and, in human cancer cell lines, arm-length aneuploidies are strongly correlated with either sensitivity or resistance to certain drugs70. Comparable results have been observed in yeast, where gaining certain chromosomes is a common mechanism by which cells can acquire resistance to antimycotics71,72. Aneuploidy may also help cells to survive some genetic perturbations; for instance, when an oncogenic driver mutation73 or a key fitness-associated gene is lost74.
Prolonged culture stress can also select for aneuploid karyotypes. While human embryonic stem cells are karyotypically stable in short-term cultures, aneuploid populations commonly arise and outcompete their euploid progenitors when these cells are grown in vitro for long periods22,75,76,77. Human embryonic stem cells commonly gain copies of chromosomes 12 and 17; these chromosomes are also frequently gained in testicular cancers that originate from pluripotent germ cells76. Thus, the fitness benefit conferred by these trisomies in vitro may recapitulate a tissue type-specific selective advantage conferred by aneuploidy during tumorigenesis22,75,76,77.
Highly aneuploid tumours are associated with metastases and poor patient outcomes
The ability of aneuploid cells to grow under suboptimal conditions may also facilitate the spread of aneuploid cancer cells to distant sites throughout the body. Indeed, metastatic lesions typically display a greater aneuploidy burden than primary tumours34,70,78,79, and aneuploidy is a biomarker of poor prognosis in a variety of cancer types, including myeloma80, prostate cancer81, melanoma82 and breast cancer83.
The relationship between aneuploidy and metastasis may be driven by aneuploidy’s ability to confer phenotypic plasticity on cancer cells. In colon cancer, gaining an extra copy of chromosome 5 slows cell division and inhibits the growth of a primary xenograft32. However, this aneuploidy also causes a partial epithelial–mesenchymal transition that triggers invasive growth and a significant increase in metastatic dissemination34. These phenotypes were found to be specific for both that individual aneuploidy and that distinct genetic background: when several other aneuploidies were introduced into the same colon cancer cell line, no epithelial–mesenchymal transition was observed, and introducing an extra copy of chromosome 5 into a different cell line failed to phenocopy its effects in colon cancer34. Aneuploidy has also been associated with cell state transitions in ovarian cancer: loss of chromosome arm 16p correlates with an epithelial–mesenchymal transition that facilitates cancer cell dissemination, while losing chromosome arm 10p correlates with a mesenchymal–epithelial transition that promotes cellular outgrowth84. Thus, chromosome copy number alterations may facilitate large-scale phenotypic changes that allow cells to grow in diverse cellular milieus, including sites of metastatic dissemination.
Mouse models reveal oncogenic and tumour-suppressive roles for CIN
Most of the experiments described in the preceding sections were conducted on cell lines manipulated to gain or lose certain chromosomes. However, genetically engineered mouse models (GEMMs) that harbour mutations that increase the rate of chromosome mis-segregation have also shed light on the relationship between aneuploidy and cancer. In general, mice with CIN display a range of protumorigenic and antitumorigenic phenotypes, underscoring a complicated relationship between CIN, aneuploidy and tumour development. For instance, some mouse studies have suggested that CIN is sufficient to initiate tumorigenesis. Mice overexpressing the spindle-assembly checkpoint component MAD2 (also known as MAD2L1) or HEC1 (also known as NDC80) display constitutive chromosome mis-segregation and a significant increase in spontaneous tumour formation49,85. However, such results are not universal across CIN models. Indeed, mice harbouring heterozygous deletions in the genes encoding the spindle-assembly checkpoint components BUB1 and BUB3 display CIN but are not predisposed to spontaneous tumour development86. Even in the MAD2-overexpressing mouse, overall tumour incidence remains relatively low, with only ~50% of mice developing tumours after a long latency period of 20 months85.
Genetically induced CIN is also capable of accelerating tumorigenesis induced by carcinogens or other genetic alterations. While Bub1-heterozygous mice are not tumour-prone on their own, treating these mice with the carcinogen 7,12-dimethylbenz[a]anthracene results in a threefold increase in tumour incidence compared with wild-type mice87. Additionally, experiments crossing mice with CIN to mice with cancer-predisposed genetic backgrounds such as Trp53+/− mice or ApcMin/+ mice reveal collaborative synergy, with an increase in tumour incidence and a decrease in tumour latency53,88,89,90. However, these synergistic effects are not universal to all cancer-predisposed genetic backgrounds, as, for example, the Bub1-heterozygous mouse does not demonstrate increased tumour incidence in an Rb1+/− or Pten+/− genetic background89. Moreover, while most models indicate that CIN potentiates tumorigenesis, some models indicate that CIN functions as a tumour suppressor. For instance, mice with heterozygous expression of either Stag1 (which encodes a cohesin subunit) or Cenpe (which encodes centromere-associated protein E) were less prone to develop tumours after carcinogen exposure when compared with wild-type mice54,91. Other studies have reported similar findings92,93,94.
To reconcile these conflicting results, it has been suggested that the degree of instability dictates whether CIN will have a tumour-promoting or tumour-suppressive effect93,95. According to this theory, low levels of CIN trigger a genomic diversification that allows evolutionary selection for growth-promoting aneuploidies, while high levels of CIN are toxic to cells and result in apoptosis (Fig. 2a,b). CIN may also have different effects on distinct stages of tumorigenesis: in the ApcMin/+ intestinal tumorigenesis model, crossing in a CIN-promoting Cenpe mutation had no effect on the formation of colon polyps but significantly inhibited their subsequent growth92. Furthermore, a recent study has highlighted how both the degree of CIN and tissue identity can affect subsequent tumour development. In this report, researchers combined hypomorphic and kinase-dead mutations in Mps1 to generate sets of mice with six different levels of CIN96, stratified according to the percentage of cells exhibiting chromosomal mis-segregation. Moderate amounts of CIN were sufficient to increase the formation of intestinal adenomas in the small intestine, while low CIN and high CIN had no effect. At the same time, in the distal part of the colon, high CIN promoted tumour formation more effectively than either low CIN or moderate CIN96. These results suggest that the cellular consequences of CIN may be distinct in different tissues, and some tissues may be able to tolerate CIN better than others (Fig. 2b).
Finally, while these GEMMs provide a proof of principle that CIN can exert both tumour-promoting and tumour-suppressive effects (Fig. 2c), the relationship between these mouse models and the development of aneuploidy during human tumorigenesis is unclear. Notably, the CIN-inducing mutations in the kinetochore, spindle-assembly checkpoint, centrosome and other mitotic complexes that have been studied in mice are extremely rare in human cancers97. Many, if not all, of the crucial genes that control CIN that have been manipulated in these GEMMs are also known to moonlight in other biological processes, including nuclear import98, transcriptional control99,100, telomere protection101 and insulin signalling102. In general, human cancer cells maintain a functional spindle-assembly checkpoint103,104, and the causes of CIN during spontaneous tumorigenesis remain debated105. Additionally, as discussed later, mis-segregating chromosomes can themselves trigger a variety of phenotypic consequences, including DNA damage, activation of the cytosolic DNA-sensing cyclic GMP–AMP synthase (cGAS)–stimulator of interferon genes (STING) pathway and chromothripsis (Box 1). Thus, these CIN models may affect tumorigenesis through both aneuploidy-dependent and aneuploidy-independent mechanisms.
Down syndrome and cancer
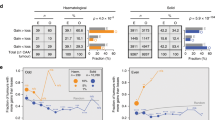
The contradictory tumour-promoting and tumour-suppressive effects of aneuploidy are clearly apparent in Down syndrome (trisomy 21), the only autosomal aneuploidy in humans that is compatible with adult survival (Fig. 2d). While advanced malignancies often display complex patterns of aneuploidy, early precancerous lesions often harbour one or a few chromosomal alterations, and thus the study of Down syndrome may shed light on the role of aneuploidy during the progression to malignant growth14,15,16,17,18,19.
Individuals with Down syndrome have an increased likelihood of developing several forms of leukaemia, including a 500-fold increased risk of acute megakaryoblastic leukaemia106,107. At the same time, individuals with Down syndrome are significantly less likely to develop most solid cancers, including lung cancer, breast cancer and cervical cancer108. Overall, individuals with Down syndrome have a 50% lower risk of non-leukaemia cancers compared with age-matched individuals without Down syndrome109. These results are particularly striking considering the fact that individuals with Down syndrome are more likely to have several risk factors for increased tumour development, including obesity110, low physical activity111 and nulliparity112.
The dual nature of Down syndrome is also apparent in the different mouse models of trisomy 21. The most commonly used model of Down syndrome is the Ts65Dn mouse, which harbours a triplication of a region of mouse chromosome 16 that is syntenic to human chromosome 21 (ref.43). Ts65Dn mice share several similarities with trisomy 21 in humans, including craniofacial abnormalities, congenital heart defects and alterations in learning and memory113,114,115,116,117,118. Additionally, the Ts65Dn triplication appears to be broadly tumour suppressive: Ts65Dn mice develop fewer lesions when crossed with ApcMin/+ mice119, exhibit slower growth of transplanted skin and lung cancer xenografts120, and survive for longer when crossed with mice that have mutations in the tumour suppressor genes Trp53 and Nf1 (ref.121). Yet, Ts65Dn mice also display alterations in haematopoiesis that resemble the transient myeloproliferative disease that precedes leukaemia development in individuals with Down syndrome122. A related Down syndrome mouse model, called ‘Tc1’, was generated by transferring a nearly complete copy of human chromosome 21 into mouse embryonic stem cells123. This model maintains both the coding sequences and the regulatory elements of the human chromosome, which may better recapitulate the consequences of Down syndrome-associated genomic imbalance. As with the Ts65Dn model, Tc1 mice are protected from the development of solid tumours but display a preleukaemic myeloproliferative disorder45,124. In both mouse models, analyses of the early stages of tumorigenesis suggest that Down syndrome blocks the development of blood vessels that support cancer growth. Genetic dissection of the triplicated regions have identified multiple genes that, when overexpressed, are capable of antagonizing tumour angiogenesis (discussed later).
In total, these studies of Down syndrome exemplify the complex relationship between aneuploidy and cancer: aneuploidy is neither universally oncogenic nor universally tumour suppressive, and the specific impact of an aneuploidy depends on both the tissue and the identity of the duplicated region. Ongoing research into Down syndrome seeks to counter the effects of aneuploidy and restore normal cell physiology, which may also have direct implications for the treatment of aneuploid cancers. At the same time, most aggressive malignancies maintain highly aneuploid karyotypes, and the single trisomy that occurs in Down syndrome and its associated mouse models may not perfectly recapitulate the phenotypes associated with complex cancer aneuploidies.
Causes of aneuploid phenotypes
Gene dosage imbalances cause many of the phenotypes observed in aneuploid cells
It is likely that the major mechanism by which aneuploidy influences tumour development is its effect on gene expression (Fig. 3). In general, aneuploidy alters transcription levels, with the average amount of mRNA produced from a chromosome scaling according to DNA copy number19,20,21,22,23,24,25. This change in transcription results in a similar and proportional change in the protein produced from an aneuploid chromosome24,125, with certain exceptions. Notably, some subunits of macromolecular complexes are subject to dosage compensation, and their protein levels may be unaffected by changes in DNA or mRNA abundance24,125,126,127. Nonetheless, across a large dataset of human cancer cell lines, the average Pearson correlation coefficient between mRNA and protein levels is ~0.5, indicating a generally strong correlation between transcription and translation128.
Aneuploidy is a consequence of chromosomal instability (CIN), but CIN can also arise due to unstable aneuploid karyotypes. Aneuploidy induces gene and protein dosage changes, while mis-segregating chromosomes can result in micronucleation, DNA damage, chromothripsis and cyclic GMP–AMP synthase (cGAS)–stimulator of interferon genes (STING) activation. Chromothripsis can have multiple effects on the cell (Box 1), including dosage changes of specific genes encoded on the double minutes and complex aneuploidy from the chromothriptic chromosome. Both extensive DNA damage and stressors caused by protein dosage changes can lead to cellular senescence, which lead to an altered secretome as cells secrete senescence-associated secretory phenotype (SASP) factors. Cytosolic DNA activates the cGAS–STING pathway, leading to cellular senescence and SASP factor secretion.
Several lines of evidence indicate the centrality of imbalanced gene expression in understanding aneuploidy-induced phenotypes. First, in general, the number of genes encoded on an aneuploid chromosome is directly proportional to the severity of the aneuploid phenotype. In humans, the three autosomal trisomies that can survive until birth affect the three chromosomes that code for the fewest genes (chromosomes 13, 18 and 21)27. All other trisomies result in early miscarriages1,2. Moreover, chromosome 21 has fewer genes than chromosomes 13 and 18, and the phenotypes of Down syndrome are much milder129 than the phenotypes of trisomy 13 or 18, which are typically lethal within the first year after birth130. A similar pattern has been observed in the mouse: while all chromosome gains and losses are embryonically lethal to mice, larger aneuploidies are lost at earlier developmental time points than aneuploidies affecting smaller chromosomes27. Secondly, an analysis of tumour suppressor genes, essential genes and proliferation-driver genes found that the locations of these genes predicted the complex aneuploidy patterns found in cancer, suggesting that the imbalanced expression of these genes resulted in the selection for these specific aneuploidies131,132. Thirdly, according to a recent preprint, aneuploidies that arise in cancer can reinforce gene expression patterns that occur in a tumour’s tissue of origin. For instance, the genes on chromosome arm 13q are highly expressed in the colon relative to other healthy tissues, and 13q is frequently gained in colon cancer133. Thus, these specific aneuploidies may be acquired to ‘lock in’ certain beneficial transcriptional programmes. Finally, in yeast, the gain of an endogenous chromosome causes multiple aneuploidy-associated phenotypes, including slow proliferation, genomic instability and metabolic alterations134. However, gaining an artificial chromosome that lacks any protein-coding sequences has almost no effect on cell physiology134. In total, these experimental results are consistent with a model in which the major driver of aneuploidy phenotypes is the imbalanced expression of genes encoded on affected chromosomes.
Aneuploidy-induced copy number changes in specific genes can both promote and suppress cancer progression. For example, expression of the oncoprotein MYC plays a critical role in acute myeloid leukaemia. In mice, MYC is encoded on chromosome 15, and approximately 60% of mice with acute myeloid leukaemia spontaneously acquire a trisomy of this chromosome135. However, when MYC expression is increased ectopically through the use of a randomly integrated transgene, only 5% of subsequent leukaemias acquire chromosome 15 trisomies135. This demonstrates that selection for trisomy 15 is driven by the phenotypic benefits conferred by MYC overexpression, and when MYC is overexpressed in an aneuploidy-independent manner, selection for this trisomy is lost.
Similarly, human chromosome 21 (and its syntenic region on mouse chromosome 16) contains multiple genes that have been demonstrated to affect tumorigenesis. Chromosome 21 harbours ERG, a gene encoding an ETS-family transcription factor that is essential for normal haematopoiesis136. Overexpression of ERG promotes the immortalization of haematopoietic progenitor cells136,137,138, while deleting a single copy of Erg blocks the development of the myeloproliferative disorder in Ts65Dn mice139. At the same time, several genes on chromosome 21 have also been suggested to function as dosage-sensitive tumour suppressors. Mouse models of Down syndrome are generally associated with a decrease in angiogenesis, and several triplicated genes, including Dscr1 (also known as Rcan1)120,140, Col18a1 (refs141,142) and Adamts1 (ref.143), have been identified that typically reduce blood vessel formation. Notably, the overexpression of a single extra copy of Dscr1 from its native promoter is sufficient to inhibit vascular endothelial growth factor (VEGF)-mediated angiogenesis and decrease the growth of transplanted xenografts120,140.
Gene dosage imbalances are also responsible for the development of many phenotypes that are shared among aneuploid cells. Aneuploidy affects protein translation on a massive scale, altering the expression of thousands of proteins simultaneously. This protein deregulation has been shown to lead to proteotoxic stress56,60,144,145,146,147, as cells struggle to cope with stoichiometric imbalances, increased demands on protein folding machinery and an accumulation of misfolded proteins24,27,58,144. This proteotoxic stress induces protein aggregation, and aneuploid cells can counteract the negative effects of this protein misfolding by inducing autophagosome-mediated protein degradation in yeast and mammalian cells144,145,148. Aneuploid mammalian cells also have an altered cellular metabolism, with increased sugar and nitrogen uptake, and enhanced production of metabolic wastes such as ammonium and lactate38,55. These metabolic alterations are likely due to the stresses and energy costs of additional translation, protein folding and protein degradation conferred by gene dosage imbalances. Finally, aneuploidy triggers replication stress, hyper-recombination and chromosome mis-segregation in both yeast and mammalian cells33,134,149. In yeast, these phenotypes are more severe in haploid aneuploidies (which harbour 1n + 1 karyotypes) compared with diploid aneuploidies (which harbour 2n + 1 karyotypes), underscoring how the magnitude of a dosage imbalance influences the subsequent physiology of aneuploid cells134.
Non-cell-autonomous effects of senescent aneuploid cells
Aneuploidy can also influence tumorigenesis through senescence. Experiments conducted in a variety of mammalian systems, including trisomic primary cells and cancer cells manipulated to exhibit high levels of CIN, have demonstrated that chromosome dosage imbalances result in cellular senescence59,150,151,152,153,154. Considered in a cell-autonomous manner, senescence is a potent mechanism of tumour suppression: an aneuploid cell that undergoes senescence will stop dividing and is unlikely to undergo cellular transformation155,156. However, senescence also results in a global change in gene expression, triggering the senescence-associated secretory phenotype (SASP). The SASP is a unique secretome of biologically active factors consisting of chemokines, cytokines, growth factors and immune regulators that are released into the microenvironment by senescent cells157,158. In a context-dependent and cell type-dependent manner, the SASP has been shown to be both tumour suppressive159,160 and protumorigenic161,162,163.
SASP secreted factors perform several important roles in senescence, including reinforcing growth arrest in neighbouring cells164 and aiding in the elimination of senescent cells via the attraction of natural killer cells165,166. Gene expression analysis of highly aneuploid cells revealed a SASP-like gene expression profile, which may contribute to the ability of the immune system to remove cells that have undergone chromosome mis-segregation167. At the same time, mixing senescent human fibroblasts with breast cancer cells promotes cell proliferation in vitro and tumour growth in nude mice168. Through the SASP, senescent cells can increase invasiveness and migration in neighbouring cells, thereby contributing to tumour progression162.
In general, the degree of aneuploidy in cancer cells correlates both with senescence and with the increased expression of proinflammatory and growth-promoting SASP factors169. In patient samples of invasive ductal breast carcinomas, higher levels of senescence, an upregulation of SASP factors and an increased incidence of abnormal ploidies were found at the invasive tumour front when compared with the tumour centre169. These findings support a paracrine-dependent, non-cell-autonomous mechanism of aneuploidy-associated tumorigenesis, in which the SASP continues to promote tumour progression, even after aneuploid-induced senescent cells may have been removed.
Mis-segregating chromosomes can have a variety of phenotypic outcomes
Aneuploidy can result from errors during the mitotic process that lead to an unequal division of a cell’s chromosomes. While in this Review we largely examine aneuploidy as a genetic phenomenon, it is nonetheless important to recognize that chromosomes are physical objects, and as such they can affect and be affected by a number of biological processes in the cell (Fig. 3). Notably, mis-segregating chromosomes can be damaged during or after mitosis, potentially leading them to shatter in a process called ‘chromothripsis’170,171,172,173 (Box 1). Furthermore, if all or part of a chromosome becomes trapped outside the nucleus following cell division, these fragments are capable of activating an inflammatory response through the cytosolic DNA-sensing cGAS–STING pathway174 (Box 2). Thus, the mechanism by which aneuploidy arises can alter its phenotypic consequences: failure of a mitotic kinesin (as in the Cenpe-heterozygous mouse model)91 may lead to polar chromosomes that escape mitosis undamaged, while centrosome amplification (as observed in the Plk4-overexpression mouse model)175 may increase the likelihood that aneuploid chromosomes are broken during cell division. As aneuploidy itself is capable of catalysing chromosome mis-segregation134, likely due to imbalances in the expression of key mitotic proteins33,134, chromothripsis, DNA damage and cGAS–STING activation may also be downstream consequences of certain forms of aneuploidy.
Aneuploidy and immune recognition
Recent evidence has demonstrated that another key cancer hallmark — the activation and evasion of immunosurveillance — can be affected by aneuploidy. According to the standard model of tumour immunosurveillance, various exogenous and endogenous stresses generate mutations in oncogenes and tumour suppressors over the course of an organism’s lifetime, pushing normal cells towards malignant transformation. These subclonal mutations also function as neoantigens, allowing most premalignant cells to be recognized and eliminated by the immune system176,177. Theoretically, aneuploidy could function as a neoantigen-independent mechanism to promote tumorigenesis: a whole-chromosome gain or loss could increase or decrease the expression of an oncogene or tumour suppressor, without actually resulting in an immunogenic change in protein sequence. Consistent with this hypothesis, highly aneuploid tumours tend to show lower levels of CD8+ T cell infiltration compared with tumours with lower levels of aneuploidy, while tumours with a high mutational load show the opposite pattern11,178. Additionally, in a retrospective analysis, aneuploidy was associated with decreased responsiveness to anti-CTLA4 immunotherapy in melanoma178. Thus, these correlative analyses suggest that aneuploidy may help tumours to escape immune-mediated destruction.
Experiments resulting from the direct manipulation of cancer cell ploidy paint a more complicated picture. In general, many of the stress-response pathways that are activated by aneuploidy are also associated with immune recognition. As discussed earlier, when CIN is induced in diploid cells with use of an MPS1 inhibitor, the highly aneuploid cells that result tend to senesce and exhibit a SASP-like gene expression profile167. In co-culture experiments, these aneuploid cells are recognized and destroyed by natural killer cells, while euploid cells are killed at a significantly lower rate167,179. Similarly, inducing ploidy changes tends to block the growth of xenografts in immunocompetent mice, while these same xenografts are capable of forming tumours in immunocompromised animals180,181. The destruction of aneuploid cells by the immune system may in part be mediated by the cGAS–STING pathway, which causes an increase in the secretion of inflammatory cytokines in response to micronucleated DNA182,183 (Fig. 4).
Aneuploidy can affect the expression of oncogenes and tumour suppressors without creating neoantigens, thereby facilitating tumorigenesis without activating immune recognition. Aneuploidy has also been shown to alter antigen and cell surface marker expression, creating a heterogeneous population of antigen and cell surface markers that allows certain clones to evade immune recognition. Chromosomal instability, an aneuploidy-associated phenotype, can activate the cyclic GMP–AMP synthase (cGAS)–stimulator of interferon genes (STING) pathway and induce the senescence-associated secretory phenotype (SASP) (Box 2). Secretion of SASP factors may allow the immune system, namely natural killer (NK) cells, to detect and eliminate aneuploid cells; however, the SASP has also been shown to play a role in protumorigenic phenotypes such as mesenchymal transitions, chronic inflammation and secretion of protumorigenic factors. ER, endoplasmic reticulum.
Recent evidence has suggested one potential way that these disparate results can be reconciled. In a colon cancer xenograft model, aneuploid tumours initially grew significantly worse in wild-type mice compared with immunocompromised mice184. However, following ‘passaging’ in immunocompetent mice, the aneuploid tumours were found to downregulate the expression of genes associated with antigen presentation. These evolved cells subsequently exhibited superior growth compared with the near-diploid parental cells, even in wild-type mice. Thus, aneuploidy may allow the selection of cancer cells with chromosome dosage imbalances that exhibit highly aggressive phenotypes that allow their escape from immunosurveillance and eventual outgrowth. More work will be required to fully reconcile the role of aneuploidy in immune evasion with its ability to trigger immune recognition.
Targeting aneuploid cells
While aneuploidy is found in the vast majority of human tumours, it is exceedingly rare in normal tissue. (Although early fluorescent in situ hybridization-based studies reported that aneuploidy was common in the liver185 and the brain186, subsequent analyses using single-cell sequencing have demonstrated that more than 95% of cells in these tissues harbour balanced karyotypes187,188). Thus, a drug or treatment that selectively eliminates aneuploid cells could be broadly useful against a variety of cancer types while exhibiting few side effects in normal tissue.
Untransformed aneuploid cells have been found to be specifically vulnerable to certain metabolic stressors in vitro. Two separate drug screens demonstrated that aneuploid cells are more sensitive than their diploid counterparts to activators of the AMP-activated protein kinase (AMPK)–peroxisome proliferator-activated receptor-γ co-activator 1α (PGC1α) axis, a central pathway that regulates energy metabolism and mitochondrial biogenesis189,190. Aneuploid cells suffer from intrinsic energy stress, and deregulating these pathways further may increase their metabolic dysregulation to toxic levels. Aneuploid cells have also been found to contain higher levels of ceramide, and further increasing the levels of ceramide is significantly more toxic to aneuploid cells compared with diploid cells191.
Another method to target aneuploid cancer cells is to exacerbate chromosome mis-segregation. As aneuploidy and CIN are tightly intertwined, increasing the rate of chromosome mis-segregation beyond tolerable levels may specifically compromise the viability of aneuploid cells192. Many aneuploid cancer cells have altered microtubule dynamics and increased microtubule–kinetochore stability; these spindle disruptions can cause improper kinetochore attachments and prevent error corrections, leading to high levels of chromosome mis-segregation193,194. In line with these observations, inhibition of the spindle-assembly checkpoint has been found to selectively kill aneuploid cancer cells by further enhancing their level of CIN195,196,197,198.
Emerging data reported in a series of preprints have also indicated that aneuploid cancer cells may be sensitive to spindle disruption via the inhibition of a mitotic kinesin, even without total checkpoint loss198,199,200. The mitotic kinesin KIF18A is generally dispensable for mammalian cell proliferation, but knockdown of KIF18A is significantly more toxic to aneuploid cells compared with diploid cells. In cells harbouring highly aneuploid karyotypes, KIF18A knockdown has been shown to alter spindle geometry and microtubule dynamics, leading to mitotic errors, micronucleus formation and a reduction in cellular viability198,199,200. These results identify KIF18A as a promising target for future drug development efforts to establish an ‘anti-aneuploidy’ therapeutic regimen.
Outlook and future directions
The Cancer Genome Atlas and other genomic surveys have made it abundantly clear that human tumours are much more than a collection of mutant oncogenes and tumour suppressors. Indeed, nearly all tumours display some degree of aneuploidy, and the frequency of specific copy number alterations rivals the frequency of mutations in key cancer genes such as KRAS and TP53 (ref.11). These same genomic analyses have also reaffirmed an observation initially made more than 40 years ago using older technologies201,202,203,204: namely, aneuploid tumours tend to be much more aggressive and are associated with significantly shorter patient survival than tumours with near-diploid karyotypes70,78,81,205. Understanding the causes and consequences of cancer aneuploidy can therefore help us to deconstruct the fundamental origins of malignant growth and potentially develop more-effective treatments.
Moving forward, we see several areas of growth for aneuploidy research. First, while GEMMs with compromised spindle-assembly checkpoints have undoubtedly shed light on the consequences of CIN, the relative scarcity of comparable mutations in human cancers may call into question the ability of these GEMMs to accurately model spontaneous tumorigenesis. Uncovering why human tumours exhibit CIN, and developing approaches to recapitulate these processes in mice, will help to elucidate the consequences of aneuploidy in vivo. ‘Standard’ mouse cancer models such as MMTV–PyMT mice and Trp53−/− mice give rise to reproducible chromosomal imbalances that resemble the aneuploidies found in human tumours206. These mouse models could potentially be co-opted to study the causes and consequences of aneuploidy in the absence of checkpoint-abrogating mutations.
Secondly, we are currently able to manipulate single genes in normal cells and in cancer much more easily than we can manipulate chromosomal aneuploidies. Whole-genome cDNA, CRISPR and RNAi libraries have made it possible to eliminate, knockdown or overexpress any individual gene in the genome and uncover the effects of these perturbations on cancer-related phenotypes131,207,208,209. We believe that improved chromosome-engineering strategies, including new microcell-mediated chromosome transfer approaches and CRISPR-mediated chromosome elimination techniques, may similarly facilitate a greater understanding of karyotype–phenotype relationships. Finally, the remarkable efficacy of immune checkpoint inhibitors that have entered routine clinical use in the past 5 years has underscored the importance of tumour–immune interactions in cancer control210. Developing new approaches to study the immunosurveillance of aneuploidy may shed light on why highly aneuploid cancers tend to display few tumour-infiltrating lymphocytes. Techniques that enhance the recruitment of immune cells to highly aneuploid tumours may be useful in combination with immune checkpoint inhibitors, thereby increasing the potency of these therapies in some of the deadliest cancers.
References
Biancotti, J. C. et al. The in vitro survival of human monosomies and trisomies as embryonic stem cells. Stem Cell Res. 9, 218–224 (2012).
van den Berg, M. M. J., van Maarle, M. C., van Wely, M. & Goddijn, M. Genetics of early miscarriage. Biochim. Biophys. Acta 1822, 1951–1959 (2012).
Weaver, B. A. & Cleveland, D. W. Does aneuploidy cause cancer? Curr. Opin. Cell Biol. 18, 658–667 (2006).
Donne, R., Saroul-Aïnama, M., Cordier, P., Celton-Morizur, S. & Desdouets, C. Polyploidy in liver development, homeostasis and disease. Nat. Rev. Gastroenterol. Hepatol. 17, 391–405 (2020).
Lens, S. M. A. & Medema, R. H. Cytokinesis defects and cancer. Nat. Rev. Cancer 19, 32–45 (2019).
Tanaka, K. et al. Tetraploidy in cancer and its possible link to aging. Cancer Sci. 109, 2632–2640 (2018).
de Smith, A. J., Trewick, A. L. & Blakemore, A. I. F. Implications of copy number variation in people with chromosomal abnormalities: potential for greater variation in copy number state may contribute to variability of phenotype. HUGO J. 4, 1–9 (2010).
Tang, Y.-C. & Amon, A. Gene copy-number alterations: a cost-benefit analysis. Cell 152, 394–405 (2013).
Hardy, P. A. & Zacharias, H. Reappraisal of the Hansemann-Boveri hypothesis on the origin of tumors. Cell Biol. Int. 29, 983–992 (2005).
Boveri, T. Concerning the origin of malignant tumours by Theodor Boveri. Translated and annotated by Henry Harris. J. Cell Sci. 121 (Suppl. 1), 1–84 (2008).
Taylor, A. M. et al. Genomic and functional approaches to understanding cancer aneuploidy. Cancer Cell 33, 676–689.e3 (2018).
Watkins, T. B. K. et al. Pervasive chromosomal instability and karyotype order in tumour evolution. Nature 587, 126–132 (2020).
Ben-David, U. & Amon, A. Context is everything: aneuploidy in cancer. Nat. Rev. Genet. 21, 44–62 (2020).
Crowell, R. E. et al. Detection of trisomy 7 in nonmalignant bronchial epithelium from lung cancer patients and individuals at risk for lung cancer. Cancer Epidemiol. Biomarkers Prev. 5, 631–637 (1996).
Aldaz, C. M., Conti, C. J., Klein-Szanto, A. J. & Slaga, T. J. Progressive dysplasia and aneuploidy are hallmarks of mouse skin papillomas: relevance to malignancy. Proc. Natl Acad. Sci. USA 84, 2029–2032 (1987).
Aldaz, C. M., Trono, D., Larcher, F., Slaga, T. J. & Conti, C. J. Sequential trisomization of chromosomes 6 and 7 in mouse skin premalignant lesions. Mol. Carcinog. 2, 22–26 (1989).
Longy, M. et al. Chromosomal analysis of colonic adenomatous polyps. Cancer Genet. Cytogenet. 49, 249–257 (1990).
Garewal, H. S., Sampliner, R., Liu, Y. & Trent, J. M. Chromosomal rearrangements in Barrett’s esophagus. A premalignant lesion of esophageal adenocarcinoma. Cancer Genet. Cytogenet. 42, 281–286 (1989).
Mian, C. et al. Fluorescence in situ hybridization in cervical smears: detection of numerical aberrations of chromosomes 7, 3, and X and relationship to HPV infection. Gynecol. Oncol. 75, 41–46 (1999).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature 578, 122–128 (2020).
Upender, M. B. et al. Chromosome transfer induced aneuploidy results in complex dysregulation of the cellular transcriptome in immortalized and cancer cells. Cancer Res. 64, 6941–6949 (2004).
Ben-David, U. et al. Aneuploidy induces profound changes in gene expression, proliferation and tumorigenicity of human pluripotent stem cells. Nat. Commun. 5, 4825 (2014). This article reveals the effects of trisomy 12, which is the most common genomic alteration in human pluripotent stem cells. Trisomy 12 causes transcriptional changes similar to germ cell tumours and increases proliferation and tumorigenicity.
FitzPatrick, D. R. Transcriptome analysis of human autosomal trisomy. Hum. Mol. Genet. 11, 3249–3256 (2002).
Stingele, S. et al. Global analysis of genome, transcriptome and proteome reveals the response to aneuploidy in human cells. Mol. Syst. Biol. 8, 608 (2012). This study performs transcriptome and proteome analysis to understand how aneuploidy alters cellular physiology. Although transcription levels of genes on extra chromosomes scale with chromosome copy number, the expression of certain proteins shows dosage compensation.
Pavelka, N. et al. Aneuploidy confers quantitative proteome changes and phenotypic variation in budding yeast. Nature 468, 321–325 (2010).
Braun, R. et al. Single chromosome aneuploidy induces genome-wide perturbation of nuclear organization and gene expression. Neoplasia 21, 401–412 (2019).
Torres, E. M., Williams, B. R. & Amon, A. Aneuploidy: cells losing their balance. Genetics 179, 737–746 (2008).
Sheltzer, J. M., Torres, E. M., Dunham, M. J. & Amon, A. Transcriptional consequences of aneuploidy. Proc. Natl Acad. Sci. USA 109, 12644–12649 (2012).
Schukken, K. M. & Foijer, F. CIN and aneuploidy: different concepts, different consequences. Bioessays 40, 1700147 (2018).
Zhu, J., Tsai, H.-J., Gordon, M. R. & Li, R. Cellular stress associated with aneuploidy. Dev. Cell 44, 420–431 (2018).
Bakhoum, S. F. & Cantley, L. C. The multifaceted role of chromosomal instability in cancer and its microenvironment. Cell 174, 1347–1360 (2018).
Sheltzer, J. M. et al. Single-chromosome gains commonly function as tumor suppressors. Cancer Cell 31, 240–255 (2017). This study reveals that many single-chromosome trisomies suppress cellular transformation. At the same time, aneuploidy may provide evolutionary flexibility, promoting the aggressive growth of tumours with complex karyotypes.
Passerini, V. et al. The presence of extra chromosomes leads to genomic instability. Nat. Commun. 7, 10754 (2016).
Vasudevan, A. et al. Single-chromosomal gains can function as metastasis suppressors and promoters in colon cancer. Dev. Cell 52, 413–428.e6 (2020).
Zhang, M. et al. Aneuploid embryonic stem cells exhibit impaired differentiation and increased neoplastic potential. EMBO J. 35, 2285–2300 (2016).
Epstein, C. J. Mouse monosomies and trisomies as experimental systems for studying mammalian aneuploidy. Trends Genet. 1, 129–134 (1985).
Hernandez, D. & Fisher, E. M. Mouse autosomal trisomy: two’s company, three’s a crowd. Trends Genet. 15, 241–247 (1999).
Williams, B. R. et al. Aneuploidy affects proliferation and spontaneous immortalization in mammalian cells. Science 322, 703–709 (2008). This study characterizes the effects of trisomies in mouse embryonic fibroblasts. These aneuploidies increase the expression of genes on the aneuploid chromosome while also impairing proliferation and causing certain metabolic alterations.
Pfau, S. J., Silberman, R. E., Knouse, K. A. & Amon, A. Aneuploidy impairs hematopoietic stem cell fitness and is selected against in regenerating tissues in vivo. Genes Dev. 30, 1395–1408 (2016).
Bolton, H. et al. Mouse model of chromosome mosaicism reveals lineage-specific depletion of aneuploid cells and normal developmental potential. Nat. Commun. 7, 11165 (2016).
Singla, S., Iwamoto-Stohl, L. K., Zhu, M. & Zernicka-Goetz, M. Autophagy-mediated apoptosis eliminates aneuploid cells in a mouse model of chromosome mosaicism. Nat. Commun. 11, 2958 (2020).
Herbst, E. W., Pluznik, D. H., Gropp, A. & Uthgennant, H. Trisomic hemopoietic stem cells of fetal origin restore hemopoiesis in lethally irradiated mice. Science 211, 1175–1177 (1981).
Davisson, M. et al. Segmental trisomy as a mouse model for down syndrome. Prog. Clin. Biol. Res. 384, 117–133 (1993).
Kazuki, Y. et al. A non-mosaic transchromosomic mouse model of down syndrome carrying the long arm of human chromosome 21. eLife 9, e56223 (2020).
Alford, K. A. et al. Perturbed hematopoiesis in the Tc1 mouse model of down syndrome. Blood 115, 2928–2937 (2010).
Dobles, M., Liberal, V., Scott, M. L., Benezra, R. & Sorger, P. K. Chromosome missegregation and apoptosis in mice lacking the mitotic checkpoint protein Mad2. Cell 101, 635–645 (2000).
Hudson, D. F. et al. Centromere protein b null mice are mitotically and meiotically normal but have lower body and testis weights. J. Cell Biol. 141, 309–319 (1998).
Howman, E. V. et al. Early disruption of centromeric chromatin organization in centromere protein A (Cenpa) null mice. Proc. Natl Acad. Sci. USA 97, 1148–1153 (2000).
Diaz-Rodríguez, E., Sotillo, R., Schvartzman, J.-M. & Benezra, R. Hec1 overexpression hyperactivates the mitotic checkpoint and induces tumor formation in vivo. Proc. Natl Acad. Sci. USA 105, 16719–16724 (2008).
Kabeche, L. & Compton, D. A. Checkpoint-independent stabilization of kinetochore-microtubule attachments by Mad2 in human cells. Curr. Biol. 22, 638–644 (2012).
Zhang, D. et al. Cre-loxP-controlled periodic Aurora-A overexpression induces mitotic abnormalities and hyperplasia in mammary glands of mouse models. Oncogene 23, 8720–8730 (2004).
Marthiens, V. et al. Centrosome amplification causes microcephaly. Nat. Cell Biol. 15, 731–740 (2013).
Mukherjee, M. et al. separase loss of function cooperates with the loss of p53 in the initiation and progression of T- and B-cell lymphoma, leukemia and aneuploidy in mice. PLoS ONE 6, (2011).
Remeseiro, S. et al. Cohesin-SA1 deficiency drives aneuploidy and tumourigenesis in mice due to impaired replication of telomeres. EMBO J. 31, 2076–2089 (2012).
Torres, E. M. et al. Effects of aneuploidy on cellular physiology and cell division in haploid yeast. Science 317, 916–924 (2007).
Sheltzer, J. M. A transcriptional and metabolic signature of primary aneuploidy is present in chromosomally unstable cancer cells and informs clinical prognosis. Cancer Res. 73, 6401–6412 (2013).
Hasaart, K. A. L. et al. Mutation accumulation and developmental lineages in normal and Down syndrome human fetal haematopoiesis. Sci. Rep. 10, 12991 (2020).
Chunduri, N. K. & Storchová, Z. The diverse consequences of aneuploidy. Nat. Cell Biol. 21, 54–62 (2019).
Andriani, G. A. et al. Whole chromosome instability induces senescence and promotes SASP. Sci. Rep. 6, 1–17 (2016). This study reveals that CIN induces DNA damage and oxidative stress that leads to premature cellular senescence in human fibroblasts. The senescent cells resulting from CIN secrete high levels of cytokines, chemokines and growth factors, characteristic of the SASP.
Torres, E. M. et al. Identification of aneuploidy-tolerating mutations. Cell 143, 71–83 (2010).
Thompson, S. L. & Compton, D. A. Proliferation of aneuploid human cells is limited by a p53-dependent mechanism. J. Cell Biol. 188, 369–381 (2010).
Soto, M. et al. p53 prohibits propagation of chromosome segregation errors that produce structural aneuploidies. Cell Rep. 19, 2423–2431 (2017).
Giam, M. et al. p53 induces senescence in the unstable progeny of aneuploid cells. Cell Cycle https://doi.org/10.1080/15384101.2020.1850968 (2019).
Li, M. et al. The ATM-p53 pathway suppresses aneuploidy-induced tumorigenesis. Proc. Natl Acad. Sci. USA 107, 14188–14193 (2010).
Simões-Sousa, S. et al. The p38α stress kinase suppresses aneuploidy tolerance by inhibiting Hif-1α. Cell Rep. 25, 749–760.e6 (2018).
López-García, C. et al. BCL9L dysfunction impairs caspase-2 expression permitting aneuploidy tolerance in colorectal cancer. Cancer Cell 31, 79–93 (2017).
Rutledge, S. D. et al. Selective advantage of trisomic human cells cultured in non-standard conditions. Sci. Rep. 6, 22828 (2016).
Lukow, D. A. et al. Chromosomal instability accelerates the evolution of resistance to anti-cancer therapies. Preprint at bioRxiv https://doi.org/10.1101/2020.09.25.314229 (2020).
Ippolito, M. R. et al. Aneuploidy-driven genome instability triggers resistance to chemotherapy. Preprint at bioRxiv https://doi.org/10.1101/2020.09.25.313924 (2020).
Shukla, A. et al. Chromosome arm aneuploidies shape tumour evolution and drug response. Nat. Commun. 11, 449 (2020).
Selmecki, A. M., Dulmage, K., Cowen, L. E., Anderson, J. B. & Berman, J. Acquisition of aneuploidy provides increased fitness during the evolution of antifungal drug resistance. PLoS Genet. 5, e1000705 (2009).
Selmecki, A., Gerami-Nejad, M., Paulson, C., Forche, A. & Berman, J. An isochromosome confers drug resistance in vivo by amplification of two genes, ERG11 and TAC1. Mol. Microbiol. 68, 624–641 (2008).
Salgueiro, L. et al. Acquisition of chromosome instability is a mechanism to evade oncogene addiction. EMBO Mol. Med. 12, e10941 (2020).
Liu, G. et al. Gene essentiality is a quantitative property linked to cellular evolvability. Cell 163, 1388–1399 (2015).
Draper, J. S. et al. Recurrent gain of chromosomes 17q and 12 in cultured human embryonic stem cells. Nat. Biotechnol. 22, 53–54 (2004).
Baker, D. E. C. et al. Adaptation to culture of human embryonic stem cells and oncogenesis in vivo. Nat. Biotechnol. 25, 207–215 (2007).
Olariu, V. et al. Modeling the evolution of culture-adapted human embryonic stem cells. Stem Cell Res. 4, 50–56 (2010).
Smith, J. C. & Sheltzer, J. M. Systematic identification of mutations and copy number alterations associated with cancer patient prognosis. eLife 7, e39217 (2018).
Bakhoum, S. F. et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature 553, 467–472 (2018).
Barlogie, B. et al. Flow cytometry in clinical cancer research. Cancer Res. 43, 3982–3997 (1983).
Stopsack, K. H. et al. Aneuploidy drives lethal progression in prostate cancer. Proc. Natl Acad. Sci. USA 116, 11390–11395 (2019).
Kheir, S. M. et al. Prognostic significance of DNA aneuploidy in stage I cutaneous melanoma. Ann. Surg. 207, 455–461 (1988).
Xu, J., Huang, L. & Li, J. DNA aneuploidy and breast cancer: a meta-analysis of 141,163 cases. Oncotarget 7, 60218–60229 (2016).
Gao, C. et al. Chromosome instability drives phenotypic switching to metastasis. Proc. Natl Acad. Sci. USA 113, 14793–14798 (2016).
Sotillo, R. et al. Mad2 overexpression promotes aneuploidy and tumorigenesis in mice. Cancer Cell 11, 9–23 (2007).
Kalitsis, P. et al. Increased chromosome instability but not cancer predisposition in haploinsufficient Bub3 mice. Genes Chromosomes Cancer 44, 29–36 (2005).
Jeganathan, K., Malureanu, L., Baker, D. J., Abraham, S. C. & van Deursen, J. M. Bub1 mediates cell death in response to chromosome missegregation and acts to suppress spontaneous tumorigenesis. J. Cell Biol. 179, 255–267 (2007).
Baker, D. J., Jin, F. & van Deursen, J. M. The yin and yang of the Cdkn2a locus in senescence and aging. Cell Cycle 7, 2795–2802 (2008).
Baker, D. J., Jin, F., Jeganathan, K. B. & van Deursen, J. M. Whole chromosome instability caused by Bub1 insufficiency drives tumorigenesis through tumor suppressor gene loss of heterozygosity. Cancer Cell 16, 475–486 (2009).
Baker, D. J., Weaver, R. L. & van Deursen, J. M. p21 both attenuates and drives senescence and aging in BubR1 progeroid mice. Cell Rep. 3, 1164–1174 (2013).
Weaver, B. A. A., Silk, A. D., Montagna, C., Verdier-Pinard, P. & Cleveland, D. W. Aneuploidy acts both oncogenically and as a tumor suppressor. Cancer Cell 11, 25–36 (2007). This article establishes that CIN has context-dependent effects on tumorigenesis depending on the methods used to induce tumour formation. While CIN increases the rates of spontaneous lymphomas and lung tumours in aged animals, it inhibits tumorigenesis resulting from certain chemical and genetic perturbations.
Zasadil, L. M. et al. High rates of chromosome missegregation suppress tumor progression but do not inhibit tumor initiation. Mol. Biol. Cell 27, 1981–1989 (2016).
Silk, A. D. et al. Chromosome missegregation rate predicts whether aneuploidy will promote or suppress tumors. Proc. Natl Acad. Sci. USA 110, E4134–E4141 (2013).
García-Higuera, I. et al. Genomic stability and tumour suppression by the APC/C cofactor Cdh1. Nat. Cell Biol. 10, 802–811 (2008).
Funk, L. C., Zasadil, L. M. & Weaver, B. A. Living in CIN: mitotic infidelity and its consequences for tumor promotion and suppression. Dev. Cell 39, 638–652 (2016).
Hoevenaar, W. H. M. et al. Degree and site of chromosomal instability define its oncogenic potential. Nat. Commun. 11, 1501 (2020). This study demonstrates that differing levels of CIN can have opposing effects in different tissues. Moderate CIN increases tumour formation in the intestine and distal part of the colon of a mouse model, but higher CIN increases tumour development in the distal part of the colon without any effect on the small intestine.
Schvartzman, J.-M., Sotillo, R. & Benezra, R. Mitotic chromosomal instability and cancer: mouse modelling of the human disease. Nat. Rev. Cancer 10, 102–115 (2010).
Wozniak, R., Burke, B. & Doye, V. Nuclear transport and the mitotic apparatus: an evolving relationship. Cell. Mol. Life Sci. 67, 2215–2230 (2010).
Balbás-Martínez, C. et al. Recurrent inactivation of STAG2 in bladder cancer is not associated with aneuploidy. Nat. Genet. 45, 1464–1469 (2013).
Solomon, D. A., Kim, J.-S. & Waldman, T. Cohesin gene mutations in tumorigenesis: from discovery to clinical significance. BMB Rep. 47, 299–310 (2014).
Cipressa, F. et al. A role for separase in telomere protection. Nat. Commun. 7, 10405 (2016).
Choi, E., Zhang, X., Xing, C. & Yu, H. Mitotic checkpoint regulators control insulin signaling and metabolic homeostasis. Cell 166, 567–581 (2016).
Tighe, A., Johnson, V. L., Albertella, M. & Taylor, S. S. Aneuploid colon cancer cells have a robust spindle checkpoint. EMBO Rep. 2, 609–614 (2001). This article demonstrates that colon cancer cell lines that display CIN nonetheless have a functional spindle-assembly checkpoint and undergo mitotic arrest on spindle damage.
Gascoigne, K. E. & Taylor, S. S. Cancer cells display profound intra- and interline variation following prolonged exposure to antimitotic drugs. Cancer Cell 14, 111–122 (2008).
Thompson, S. L., Bakhoum, S. F. & Compton, D. A. Mechanisms of chromosomal instability. Curr. Biol. 20, R285–R295 (2010).
Hasle, H., Clemmensen, I. H. & Mikkelsen, M. Risks of leukaemia and solid tumours in individuals with Down’s syndrome. Lancet 355, 165–169 (2000).
Nižetić, D. & Groet, J. Tumorigenesis in Down’s syndrome: big lessons from a small chromosome. Nat. Rev. Cancer 12, 721–732 (2012).
Hasle, H., Friedman, J. M., Olsen, J. H. & Rasmussen, S. A. Low risk of solid tumors in persons with Down syndrome. Genet. Med. 18, 1151–1157 (2016).
Patja, K., Pukkala, E., Sund, R., Iivanainen, M. & Kaski, M. Cancer incidence of persons with down syndrome in Finland: a population-based study. Int. J. Cancer 118, 1769–1772 (2006).
Nixon, D. W. Down syndrome, obesity, Alzheimer’s disease, and cancer: a brief review and hypothesis. Brain Sci. 8, 53 (2018).
Mendonca, G. V., Pereira, F. D. & Fernhall, B. Reduced exercise capacity in persons with Down syndrome: cause, effect, and management. Ther. Clin. Risk Manag. 6, 601–610 (2010).
Patja, K., Eero, P. & Iivanainen, M. Cancer incidence among people with intellectual disability. J. Intellect. Disabil. Res. 45, 300–307 (2001).
Demas, G. E., Nelson, R. J., Krueger, B. K. & Yarowsky, P. J. Spatial memory deficits in segmental trisomic Ts65Dn mice. Behav. Brain Res. 82, 85–92 (1996).
Baxter, L. L., Moran, T. H., Richtsmeier, J. T., Troncoso, J. & Reeves, R. H. Discovery and genetic localization of Down syndrome cerebellar phenotypes using the Ts65Dn mouse. Hum. Mol. Genet. 9, 195–202 (2000).
Richtsmeier, J. T., Baxter, L. L. & Reeves, R. H. Parallels of craniofacial maldevelopment in Down syndrome and Ts65Dn mice. Dev. Dyn. 217, 137–145 (2000).
Belichenko, P. V. et al. Synaptic structural abnormalities in the Ts65Dn mouse model of Down Syndrome. J. Comp. Neurol. 480, 281–298 (2004).
Kleschevnikov, A. M. et al. Hippocampal long-term potentiation suppressed by increased inhibition in the Ts65Dn mouse, a genetic model of Down syndrome. J. Neurosci. 24, 8153–8160 (2004).
Moore, C. S. Postnatal lethality and cardiac anomalies in the Ts65Dn Down syndrome mouse model. Mamm. Genome 17, 1005–1012 (2006).
Sussan, T. E., Yang, A., Li, F., Ostrowski, M. C. & Reeves, R. H. Trisomy represses ApcMin-mediated tumours in mouse models of Down’s syndrome. Nature 451, 73–75 (2008).
Baek, K.-H. et al. Down’s syndrome suppression of tumour growth and the role of the calcineurin inhibitor DSCR1. Nature 459, 1126–1130 (2009).
Yang, A. & Reeves, R. H. Increased survival following tumorigenesis in Ts65Dn mice that model Down syndrome. Cancer Res. 71, 3573–3581 (2011).
Kirsammer, G. et al. Highly penetrant myeloproliferative disease in the Ts65Dn mouse model of Down syndrome. Blood 111, 767–775 (2008).
O’Doherty, A. et al. An aneuploid mouse strain carrying human chromosome 21 with Down syndrome phenotypes. Science 309, 2033–2037 (2005).
Reynolds, L. E. et al. Tumour angiogenesis is reduced in the Tc1 mouse model of Down syndrome. Nature 465, 813–817 (2010). This article reveals that transplanted tumour cells grow poorly when injected into Tc1 mice due to decreased angiogenesis. Depletion of putative antiangiogenic genes such as Adamts1 and Erg from the extra copy of chromosome 21 is sufficient to restore a normal angiogenic response.
Nusinow, D. P. et al. Quantitative proteomics of the Cancer Cell Line Encyclopedia. Cell 180, 387–402.e16 (2020).
Dephoure, N. et al. Quantitative proteomic analysis reveals posttranslational responses to aneuploidy in yeast. eLife 3, e03023 (2014).
McShane, E. et al. Kinetic analysis of protein stability reveals age-dependent degradation. Cell 167, 803–815.e21 (2016).
Gry, M. et al. Correlations between RNA and protein expression profiles in 23 human cell lines. BMC Genomics 10, 365 (2009).
Roizen, N. J. & Patterson, D. Down’s syndrome. Lancet 361, 1281–1289 (2003).
Wu, J., Springett, A. & Morris, J. K. Survival of trisomy 18 (Edwards syndrome) and trisomy 13 (Patau syndrome) in England and Wales: 2004-2011. Am. J. Med. Genet. A 161A, 2512–2518 (2013).
Sack, L. M. et al. Profound tissue specificity in proliferation control underlies cancer drivers and aneuploidy patterns. Cell 173, 499–514.e23 (2018).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013). This study develops novel computational models to analyse mutational signatures during human tumorigenesis to classify genes as oncogenes or tumour suppressors. The distribution of these genes on chromosomes correlates with the complex patterns of copy number alterations found in cancer genomes.
Auslander, N. et al. Cancer-type specific aneuploidies hard-wire chromosome-wide gene expression patterns of their tissue of origin. Preprint at bioRxiv https://doi.org/10.1101/563858 (2019).
Sheltzer, J. M. et al. Aneuploidy drives genomic instability in yeast. Science 333, 1026–1030 (2011).
Jones, L. et al. Gain of MYC underlies recurrent trisomy of the MYC chromosome in acute promyelocytic leukemia. J. Exp. Med. 207, 2581–2594 (2010). This article demonstrates that the gain of mouse chromosome 15, harbouring the Myc oncogene, occurs spontaneously during the development of acute myeloid leukaemia. In a mouse model, the selection of trisomy 15 is due to the fitness advantages provided by MYC overexpression, and when MYC is overexpressed ectopically, selection for this trisomy is lost.
Tursky, M. L. et al. Overexpression of ERG in cord blood progenitors promotes expansion and recapitulates molecular signatures of high ERG leukemias. Leukemia 29, 819–827 (2015).
Carmichael, C. L. et al. Hematopoietic overexpression of the transcription factor Erg induces lymphoid and erythro-megakaryocytic leukemia. Proc. Natl Acad. Sci. USA 109, 15437–15442 (2012).
Stankiewicz, M. J. & Crispino, J. D. ETS2 and ERG promote megakaryopoiesis and synergize with alterations in GATA-1 to immortalize hematopoietic progenitor cells. Blood 113, 3337–3347 (2009).
Ng, A. P. et al. Trisomy of Erg is required for myeloproliferation in a mouse model of Down syndrome. Blood 115, 3966–3969 (2010). This article identifies a gene on chromosome 21 that can contribute to the haematopoietic disorders found in individuals with Down syndrome. The authors demonstrate that losing a single copy of the Erg gene reverses the pathological and haematological features of myeloproliferation such as megakaryocytosis and progenitor cell expansion in Ts65Dn mice.
Shin, J., Lee, J. C. & Baek, K.-H. A single extra copy of Dscr1 improves survival of mice developing spontaneous lung tumors through suppression of tumor angiogenesis. Cancer Lett. 342, 70–81 (2014).
O’Reilly, M. S. et al. Endostatin: an endogenous inhibitor of angiogenesis and tumor growth. Cell 88, 277–285 (1997).
Zorick, T. S. et al. High serum endostatin levels in down syndrome: implications for improved treatment and prevention of solid tumours. Eur. J. Hum. Genet. 9, 811–814 (2001).
Obika, M. et al. Tumor growth inhibitory effect of ADAMTS1 is accompanied by the inhibition of tumor angiogenesis. Cancer Sci. 103, 1889–1897 (2012).
Oromendia, A. B., Dodgson, S. E. & Amon, A. Aneuploidy causes proteotoxic stress in yeast. Genes Dev. 26, 2696–2708 (2012).
Santaguida, S., Vasile, E., White, E. & Amon, A. Aneuploidy-induced cellular stresses limit autophagic degradation. Genes Dev. 29, 2010–2021 (2015). This article shows that aneuploid cells undergo a lysosomal stress response due to an increase in protein aggregation. This proteotoxic stress activates the transcription factor TFEB to upregulate the expression of genes required for autophagy-mediated protein degradation.
Dodgson, S. E. et al. Chromosome-specific and global effects of aneuploidy in Saccharomyces cerevisiae. Genetics 202, 1395–1409 (2016).
Santaguida, S. & Amon, A. Aneuploidy triggers a TFEB-mediated lysosomal stress response. Autophagy 11, 2383–2384 (2015).
Donnelly, N., Passerini, V., Dürrbaum, M., Stingele, S. & Storchová, Z. HSF1 deficiency and impaired HSP90-dependent protein folding are hallmarks of aneuploid human cells. EMBO J. 33, 2374–2387 (2014).
Nicholson, J. M. et al. Chromosome mis-segregation and cytokinesis failure in trisomic human cells. eLife 4, e05068 (2015).
Lentini, L., Barra, V., Schillaci, T. & Di Leonardo, A. MAD2 depletion triggers premature cellular senescence in human primary fibroblasts by activating a p53 pathway preventing aneuploid cells propagation. J. Cell. Physiol. 227, 3324–3332 (2012).
Humbert, N. et al. Regulation of ploidy and senescence by the AMPK-related kinase NUAK1. EMBO J. 29, 376–386 (2010).
Meena, J. K. et al. Telomerase abrogates aneuploidy-induced telomere replication stress, senescence and cell depletion. EMBO J. 34, 1371–1384 (2015).
Macedo, J. C. et al. FoxM1 repression during human aging leads to mitotic decline and aneuploidy-driven full senescence. Nat. Commun. 9, 2834 (2018).
Barroso-Vilares, M. et al. Small-molecule inhibition of aging-associated chromosomal instability delays cellular senescence. EMBO Rep. 21, e49248 (2020).
Collado, M. & Serrano, M. Senescence in tumours: evidence from mice and humans. Nat. Rev. Cancer 10, 51–57 (2010).
Prieur, A. & Peeper, D. S. Cellular senescence in vivo: a barrier to tumorigenesis. Curr. Opin. Cell Biol. 20, 150–155 (2008).
Freund, A., Orjalo, A. V., Desprez, P.-Y. & Campisi, J. Inflammatory networks during cellular senescence: causes and consequences. Trends Mol. Med. 16, 238–246 (2010).
Pawlikowski, J. S., Adams, P. D. & Nelson, D. M. Senescence at a glance. J. Cell Sci. 126, 4061–4067 (2013).
Serrano, M. Final act of senescence. Nature 479, 481–482 (2011).
Hoenicke, L. & Zender, L. Immune surveillance of senescent cells — biological significance in cancer- and non-cancer pathologies. Carcinogenesis 33, 1123–1126 (2012).
Haugstetter, A. M. et al. Cellular senescence predicts treatment outcome in metastasised colorectal cancer. Br. J. Cancer 103, 505–509 (2010).
Coppé, J.-P. et al. Senescence-associated secretory phenotypes reveal cell-nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol. 6, 2853–2868 (2008).
Rodier, F. et al. Persistent DNA damage signaling triggers senescence-associated inflammatory cytokine secretion. Nat. Cell Biol. 11, 973–979 (2009).
Acosta, J. C. et al. Chemokine signaling via the CXCR2 receptor reinforces senescence. Cell 133, 1006–1018 (2008).
Kuilman, T. et al. Oncogene-induced senescence relayed by an interleukin-dependent inflammatory network. Cell 133, 1019–1031 (2008).
Raulet, D. H. & Guerra, N. Oncogenic stress sensed by the immune system: role of NK cell receptors. Nat. Rev. Immunol. 9, 568–580 (2009).
Santaguida, S. et al. Chromosome mis-segregation generates cell-cycle-arrested cells with complex karyotypes that are eliminated by the immune system. Dev. Cell 41, 638–651.e5 (2017). This study describes a mechanism by which the immune system can detect and eliminate aneuploid cells. Aneuploid cells exhibit senescent features and secrete proinflammatory signals, which can promote their own immune clearance.
Krtolica, A., Parrinello, S., Lockett, S., Desprez, P. Y. & Campisi, J. Senescent fibroblasts promote epithelial cell growth and tumorigenesis: a link between cancer and aging. Proc. Natl Acad. Sci. USA 98, 12072–12077 (2001).
He, Q. et al. Chromosomal instability-induced senescence potentiates cell non-autonomous tumourigenic effects. Oncogenesis 7, 62 (2018).
Cortés-Ciriano, I. et al. Comprehensive analysis of chromothripsis in 2,658 human cancers using whole-genome sequencing. Nat. Genet. 52, 331–341 (2020).
Kneissig, M. et al. Micronuclei-based model system reveals functional consequences of chromothripsis in human cells. eLife 8, e50292 (2019).
Ly, P. & Cleveland, D. W. Rebuilding chromosomes after catastrophe: emerging mechanisms of chromothripsis. Trends Cell Biol. 27, 917–930 (2017).
Forment, J. V., Kaidi, A. & Jackson, S. P. Chromothripsis and cancer: causes and consequences of chromosome shattering. Nat. Rev. Cancer 12, 663–670 (2012).
Hong, C., Tijhuis, A. E. & Foijer, F. The cGAS paradox: contrasting roles for cGAS-STING pathway in chromosomal instability. Cells 8, 1228 (2019).
Levine, M. S. et al. Centrosome amplification is sufficient to promote spontaneous tumorigenesis in mammals. Dev. Cell 40, 313–322.e5 (2017).
Yarchoan, M., Johnson, B. A., Lutz, E. R., Laheru, D. A. & Jaffee, E. M. Targeting neoantigens to augment antitumour immunity. Nat. Rev. Cancer 17, 569 (2017).
Swann, J. B. & Smyth, M. J. Immune surveillance of tumors. J. Clin. Invest. 117, 1137–1146 (2007).
Davoli, T., Uno, H., Wooten, E. C. & Elledge, S. J. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science 355, eaaf8399 (2017).
Wang, R. W., Viganò, S., Ben-David, U., Amon, A. & Santaguida, S. Aneuploid cells activate NF-κB to promote their immune clearance by NK cells. Preprint at bioRxiv https://doi.org/10.1101/2020.06.25.172239 (2020).
Senovilla, L. et al. An immunosurveillance mechanism controls cancer cell ploidy. Science 337, 1678–1684 (2012).
Boilève, A. et al. Immunosurveillance against tetraploidization-induced colon tumorigenesis. Cell Cycle 12, 473–479 (2013).
Mackenzie, K. J. et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature 548, 461–465 (2017). This study reveals how damaged DNA is detected in the cytosol by cGAS, leading to an inflammatory response. The detection of mis-segregated DNA in micronuclei by cGAS could serve as a cell-intrinsic immunosurveillance programme to clear potentially tumorigenic cells.
Deng, L. et al. STING-dependent cytosolic DNA sensing promotes radiation-induced type I interferon-dependent antitumor immunity in immunogenic tumors. Immunity 41, 843–852 (2014).
Tripathi, R., Modur, V., Senovilla, L., Kroemer, G. & Komurov, K. Suppression of tumor antigen presentation during aneuploid tumor evolution contributes to immune evasion. Oncoimmunology 8, 1657374 (2019).
Duncan, A. W. et al. Frequent aneuploidy among normal human hepatocytes. Gastroenterology 142, 25–28 (2012).
Pack, S. D. et al. Individual adult human neurons display aneuploidy: detection by fluorescence in situ hybridization and single neuron PCR. Cell Cycle 4, 1758–1760 (2005).
Knouse, K. A., Wu, J., Whittaker, C. A. & Amon, A. Single cell sequencing reveals low levels of aneuploidy across mammalian tissues. Proc. Natl Acad. Sci. USA 111, 13409–13414 (2014).
van den Bos, H. et al. Single-cell whole genome sequencing reveals no evidence for common aneuploidy in normal and Alzheimer’s disease neurons. Genome Biol. 17, 116 (2016). Knouse et al. (2014) and van den Bos et al. (2016) use single-cell sequencing to demonstrate that aneuploidy is rare in normal somatic tissue and in neurons from individuals with Alzheimer disease. These findings refute previous studies that reported high levels of aneuploidy in the brain and liver and that linked aneuploidy to the pathogenesis of Alzheimer disease.
Tang, Y.-C., Williams, B. R., Siegel, J. J. & Amon, A. Identification of aneuploidy-selective antiproliferation compounds. Cell 144, 499–512 (2011).
Schukken, K. M. et al. Altering microtubule dynamics is synergistically toxic with spindle assembly checkpoint inhibition. Life Sci. Alliance 3, e201900499 (2020).
Tang, Y.-C. et al. Aneuploid cell survival relies upon sphingolipid homeostasis. Cancer Res. 77, 5272–5286 (2017).
Janssen, A., Kops, G. J. P. L. & Medema, R. H. Elevating the frequency of chromosome mis-segregation as a strategy to kill tumor cells. Proc. Natl Acad. Sci. USA 106, 19108–19113 (2009).
Bakhoum, S. F., Thompson, S. L., Manning, A. L. & Compton, D. A. Genome stability is ensured by temporal control of kinetochore-microtubule dynamics. Nat. Cell Biol. 11, 27–35 (2009).
Ertych, N. et al. Increased microtubule assembly rates influence chromosomal instability in colorectal cancer cells. Nat. Cell Biol. 16, 779–791 (2014).
Maia, A. R. R. et al. Inhibition of the spindle assembly checkpoint kinase TTK enhances the efficacy of docetaxel in a triple-negative breast cancer model. Ann. Oncol. 26, 2180–2192 (2015).
Kops, G. J. P. L., Foltz, D. R. & Cleveland, D. W. Lethality to human cancer cells through massive chromosome loss by inhibition of the mitotic checkpoint. Proc. Natl Acad. Sci. USA 101, 8699–8704 (2004).
Wengner, A. M. et al. Novel Mps1 kinase inhibitors with potent antitumor activity. Mol. Cancer Ther. 15, 583–592 (2016).
Cohen-Sharir, Y. et al. Selective vulnerability of aneuploid human cancer cells to inhibition of the spindle assembly checkpoint. Preprint at bioRxiv https://doi.org/10.1101/2020.06.18.159038 (2020).
Quinton, R. J. et al. Whole genome doubling confers unique genetic vulnerabilities on tumor cells. Preprint at bioRxiv https://doi.org/10.1101/2020.06.18.159095 (2020).
Marquis, C. et al. Chromosomally unstable tumor cells specifically require KIF18A for proliferation. Preprint at bioRxiv https://doi.org/10.1101/2020.06.18.159327 (2020). The preprints Cohen-Sharir et al. (2020), Quinton et al. (2020) and Marquis et al. (2020) demonstrate that highly aneuploid cells are dependent on the kinesin KIF18A for mitotic progression. KIF18A inhibition may therefore serve as a strategy to selectively target aneuploid tumours.
Falor, W. H. & Ward, R. M. Prognosis in early carcinoma of the bladder based on chromosomal analysis. J. Urol. 119, 44–48 (1978).
Alimena, G., Annino, L., Balestrazzi, P., Montuoro, A. & Dallapiccola, B. Cytogenetic studies in acute leukaemias. Prognostic implications of chromosome imbalances. Acta Haematol. 58, 234–239 (1977).
Zetterberg, A. & Esposti, P. L. Prognostic significance of nuclear DNA levels in prostatic carcinoma. Scand. J. Urol. Nephrol. Suppl. 55, 53–58 (1980).
Okagaki, T., Meyer, A. A. & Sciarra, J. J. Prognosis of irradiated carcinoma of cervix uteri and nuclear DNA in cytologic postirradiation dysplasia. Cancer 33, 647–652 (1974).
Hieronymus, H. et al. Tumor copy number alteration burden is a pan-cancer prognostic factor associated with recurrence and death. eLife 7, e37294 (2018).
Ben-David, U. et al. The landscape of chromosomal aberrations in breast cancer mouse models reveals driver-specific routes to tumorigenesis. Nat. Commun. 7, 12160 (2016).
Sanson, K. R. et al. Optimized libraries for CRISPR-Cas9 genetic screens with multiple modalities. Nat. Commun. 9, 5416 (2018).
Zhan, T., Rindtorff, N., Betge, J., Ebert, M. P. & Boutros, M. CRISPR/Cas9 for cancer research and therapy. Semin. Cancer Biol. 55, 106–119 (2019).
Lin, A. & Sheltzer, J. M. Discovering and validating cancer genetic dependencies: approaches and pitfalls. Nat. Rev. Genet. 21, 671–682 (2020).
Esfahani, K. et al. A review of cancer immunotherapy: from the past, to the present, to the future. Curr. Oncol. 27, S87–S97 (2020).
Fournier, R. E. A general high-efficiency procedure for production of microcell hybrids. Proc. Natl Acad. Sci. USA 78, 6349–6353 (1981).
Bennett, A. et al. Cenp-E inhibitor GSK923295: Novel synthetic route and use as a tool to generate aneuploidy. Oncotarget 6, 20921–20932 (2015).
Thomas, R., Marks, D. H., Chin, Y. & Benezra, R. Whole chromosome loss and associated breakage-fusion-bridge cycles transform mouse tetraploid cells. EMBO J. 37, 201–218 (2018).
Kimura, M. et al. Proliferation dynamics in cultured skin fibroblasts from Down syndrome subjects. Free Radic. Biol. Med. 39, 374–380 (2005).
Hwang, S. et al. Suppressing aneuploidy-associated phenotypes improves the fitness of trisomy 21 cells. Cell Rep. 29, 2473–2488.e5 (2019).
Zuo, E. et al. CRISPR/Cas9-mediated targeted chromosome elimination. Genome Biol. 18, 224 (2017).
Giuliano, C. J., Lin, A., Girish, V. & Sheltzer, J. M. Generating single cell–derived knockout clones in mammalian cells with CRISPR/Cas9. Curr. Protoc. Mol. Biol. 128, e100 (2019).
Fernández-Miranda, G. et al. Genetic disruption of aurora B uncovers an essential role for aurora C during early mammalian development. Dev. Camb. Engl. 138, 2661–2672 (2011).
Ricke, R. M., Jeganathan, K. B. & van Deursen, J. M. Bub1 overexpression induces aneuploidy and tumor formation through Aurora B kinase hyperactivation. J. Cell Biol. 193, 1049–1064 (2011).
Rao, C. V. et al. Colonic tumorigenesis in BubR1+/–ApcMin/+ compound mutant mice is linked to premature separation of sister chromatids and enhanced genomic instability. Proc. Natl Acad. Sci. USA 102, 4365–4370 (2005).
Dai, W. et al. Slippage of mitotic arrest and enhanced tumor development in mice with BubR1 haploinsufficiency. Cancer Res. 64, 440–445 (2004).
Baker, D. J. et al. Opposing roles for p16Ink4a and p19Arf in senescence and ageing caused by BubR1 insufficiency. Nat. Cell Biol. 10, 825–836 (2008).
Nam, H.-J. & van Deursen, J. M. Cyclin B2 and p53 control proper timing of centrosome separation. Nat. Cell Biol. 16, 538–549 (2014).
Li, M., Fang, X., Wei, Z., York, J. P. & Zhang, P. Loss of spindle assembly checkpoint–mediated inhibition of Cdc20 promotes tumorigenesis in mice. J. Cell Biol. 185, 983–994 (2009).
Mukherjee, M. et al. MMTV-Espl1 transgenic mice develop aneuploid, estrogen receptor alpha (ERα)-positive mammary adenocarcinomas. Oncogene 33, 5511–5522 (2014).
Iwanaga, Y. et al. Heterozygous deletion of mitotic arrest-deficient protein 1 (MAD1) increases the incidence of tumors in mice. Cancer Res. 67, 160–166 (2007).
Michel, L. S. et al. MAD2 haplo-insufficiency causes premature anaphase and chromosome instability in mammalian cells. Nature 409, 355–359 (2001).
Chi, Y.-H., Ward, J. M., Cheng, L. I., Yasunaga, J. & Jeang, K.-T. Spindle assembly checkpoint and p53 deficiencies cooperate for tumorigenesis in mice. Int. J. Cancer 124, 1483–1489 (2009).
Foijer, F. et al. Deletion of the MAD2L1 spindle assembly checkpoint gene is tolerated in mouse models of acute T-cell lymphoma and hepatocellular carcinoma. eLife 6, e20873 (2017).
Foijer, F. et al. Chromosome instability induced by Mps1 and p53 mutation generates aggressive lymphomas exhibiting aneuploidy-induced stress. Proc. Natl Acad. Sci. USA 111, 13427–13432 (2014).
Lu, L.-Y. et al. Polo-like kinase 1 is essential for early embryonic development and tumor suppression. Mol. Cell. Biol. 28, 6870–6876 (2008).
Ko, M. A. et al. Plk4 haploinsufficiency causes mitotic infidelity and carcinogenesis. Nat. Genet. 37, 883–888 (2005).
Rosario, C. O. et al. Plk4 is required for cytokinesis and maintenance of chromosomal stability. Proc. Natl Acad. Sci. USA 107, 6888–6893 (2010).
Coelho, P. A. et al. Over-expression of Plk4 induces centrosome amplification, loss of primary cilia and associated tissue hyperplasia in the mouse. Open Biol. 5, 150209 (2015).
Babu, J. R. et al. Rae1 is an essential mitotic checkpoint regulator that cooperates with Bub3 to prevent chromosome missegregation. J. Cell Biol. 160, 341–353 (2003).
Aguirre-Portolés, C. et al. Tpx2 controls spindle integrity, genome stability, and tumor development. Cancer Res. 72, 1518–1528 (2012).
van Ree, J. H., Jeganathan, K. B., Malureanu, L. & van Deursen, J. M. Overexpression of the E2 ubiquitin–conjugating enzyme UbcH10 causes chromosome missegregation and tumor formation. J. Cell Biol. 188, 83–100 (2010).
Zhang, C.-Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015). This report demonstrates that chromothripsis can arise as a result of chromosome mis-segregation. The authors show that mis-segregated chromosomes can become trapped in micronuclei, where the DNA is subjected to genomic damage and may undergo a series of chromothriptic rearrangements.
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Huang, Y., Fenech, M. & Shi, Q. Micronucleus formation detected by live-cell imaging. Mutagenesis 26, 133–138 (2011).
Hatch, E. M., Fischer, A. H., Deerinck, T. J. & Hetzer, M. W. Catastrophic nuclear envelope collapse in cancer cell micronuclei. Cell 154, 47–60 (2013).
Storlazzi, C. T. et al. Gene amplification as double minutes or homogeneously staining regions in solid tumors: Origin and structure. Genome Res. 20, 1198–1206 (2010).
de Oliveira Mann, C. C. & Kranzusch, P. J. cGAS conducts micronuclei DNA surveillance. Trends Cell Biol. 27, 697–698 (2017).
Ohtani, N. Deciphering the mechanism for induction of senescence-associated secretory phenotype (SASP) and its role in ageing and cancer development. J. Biochem. 166, 289–295 (2019).
Zierhut, C. et al. The cytoplasmic DNA sensor cGAS promotes mitotic cell death. Cell 178, 302–315.e23 (2019).
Acknowledgements
The authors thank members of the Sheltzer laboratory for helpful comments on the manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to all aspects of the article (researching data, substantially contributing to discussion of content, and writing, reviewing and editing the manuscript before submission).
Corresponding author
Ethics declarations
Competing interests
J.M.S. is a co-founder of Meliora Therapeutics, is a member of the Scientific Advisory Board of Tyra Biosciences and has received consulting fees from Merck and Ono Pharmaceutical Co. The other authors declare no competing interests.
Additional information
J.M.S. dedicates this article to the memory of Angelika Amon, for her outstanding mentorship, her passion for tackling unorthodox questions and her pioneering research into the consequences of aneuploidy.
Peer review information
Nature Reviews Cancer thanks A. Holland, G. Kops and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Polyploidy
-
Describing an abnormal number of chromosomes, such that a cell’s karyotype is an integer multiple of the haploid complement greater than 2.
- Chromosomal instability
-
(CIN). An abnormally high rate of chromosomal mis-segregation that may result in aneuploidy.
- Microcell-mediated chromosome transfer
-
Technique involving the transfer of specific donor chromosomes into recipient cell lines as a means of creating aneuploid cells.
- Spindle-assembly checkpoint
-
Cell cycle checkpoint that maintains genome stability by ensuring proper chromosome attachment at the kinetochore via anchorage to the microtubule spindle apparatus before the initiation of anaphase.
- Cre–lox-mediated recombination
-
A technique used to introduce site-specific deletions, insertions, translocations and inversions within chromosomes.
- Robertsonian translocations
-
Translocations in which two acrocentric chromosomes fuse to share a single centromere.
- Proteotoxic stress
-
Impaired cell function arising as a result of defective protein translation, folding and/or turnover.
- Epithelial–mesenchymal transition
-
The process by which epithelial cells transition into mesenchymal cells through the loss of cell polarity and cell–cell adhesion and the subsequent gain of migratory and invasive abilities.
- Mesenchymal–epithelial transition
-
The process by which mesenchymal cells lose their migratory and invasive properties and transition into epithelial cells capable of increased cell–cell adhesion.
- Apc Min/+ mice
-
Mice that develop spontaneous intestinal adenomas due to a point mutation in the mouse homologue of the APC gene. They are frequently used as models for intestinal tumorigenesis.
- Hypomorphic
-
A mutation resulting in the reduction, but not the complete loss, of gene function.
- Cyclic GMP–AMP synthase (cGAS)–stimulator of interferon genes (STING)
-
Immune system pathway that detects cytosolic DNA and elicits a downstream inflammatory response to aid in immune clearance.
- Down syndrome
-
Genetic disorder associated with developmental and intellectual delays due to the added presence of a third copy of chromosome 21.
- Ts65Dn mouse
-
A commonly used genetic mouse model for Down syndrome that is segmentally trisomic for part of mouse chromosome 16, which encodes genes homologous to those found on human chromosome 21.
- Pearson correlation coefficient
-
Statistic measuring the strength of association between two variables.
- Replication stress
-
A cellular state in which the fidelity of DNA replication is compromised, often characterized by the frequent collapse of the replication fork.
- Hyper-recombination
-
A cellular state defined by elevated levels of recombination.
- Senescence
-
Process by which proliferating cells cease dividing due to extracellular or intracellular stress and enter a state of permanent cell cycle arrest.
- Senescence-associated secretory phenotype
-
(SASP). Unique secretome consisting of chemokines, cytokines, growth factors and immune regulators that are released into the microenvironment by senescent cells.
- MMTV–PyMT mice
-
Common model of metastatic breast cancer in which the mouse mammary tumour virus (MMTV) promotor drives the expression of the polyomavirus middle T antigen (PyMT), leading to spontaneous and rapid tumorigenesis in mouse mammary epithelial cells.
Rights and permissions
About this article
Cite this article
Vasudevan, A., Schukken, K.M., Sausville, E.L. et al. Aneuploidy as a promoter and suppressor of malignant growth. Nat Rev Cancer 21, 89–103 (2021). https://doi.org/10.1038/s41568-020-00321-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-020-00321-1
This article is cited by
-
Aneuploid serves as a prognostic marker and favors immunosuppressive microenvironment in ovarian cancer
Journal of Ovarian Research (2024)
-
PRAME induces genomic instability in uveal melanoma
Oncogene (2024)
-
Evolving copy number gains promote tumor expansion and bolster mutational diversification
Nature Communications (2024)
-
The two sides of chromosomal instability: drivers and brakes in cancer
Signal Transduction and Targeted Therapy (2024)
-
Aneuploid embryonic stem cells drive teratoma metastasis
Nature Communications (2024)